Abstract
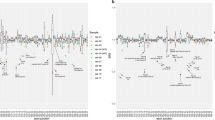
The neurofibromatosis type 1 (NF1) gene located at 17q 11.2 contains 60 exons and spans 350 kb of genomic DNA. Mutation analysis has been hampered by the large size of the gene, the high rate of new mutations, a lack of mutational clustering and the presence of numerous homologous loci. Mutation detection methods based on the direct analysis of a gene's RNA transcript permit the rapid screening of large multi-exonic genes. However, the detection of frame-shift or nonsense mutations can be limited by instability of the mutant mRNA species due to nonsense-mediated decay. In order to determine the frequency of this allelic exclusion, total lymphocyte RNA was analysed from 15 NF1 patients with known truncating mutations and a panel of 40 NF1 patients with unknown mutations. The level of expression of the mutant message was greatly reduced in 2 of the 15 samples (13%), and 3 of the 18 informative samples from the panel of 40. A coupled reverse-transcription polymerase chain reaction and protein truncation test method was subsequently applied to screen RNA from the panel of 40 unrelated NF1 patients. Aberrant polypeptide bands were identified and characterised in 21 samples (53%). The mutations identified were 479del107;ins31, 495delTGTT, 1127delTGAT, R416X, R440X, 1446del 62, 1541delAG, 2252del 74, 2537insTG, 3456delACTC, R1276X, R1362X, 5749ins171, 6084del280, 6487insA, R2214X, 6791insA, 6858del141, 7458delC, 7676 2A-G and 8081delC. These mutations were uniformly distributed across the gene and 14 represent novel changes that contribute to the germline mutational spectrum of the NF1 gene.
Similar content being viewed by others
Author information
Authors and Affiliations
Additional information
Electronic Publication
Rights and permissions
About this article
Cite this article
Osborn, M., Upadhyaya, M. Evaluation of the protein truncation test and mutation detection in the NF1 gene: mutational analysis of 15 known and 40 unknown mutations. Hum Genet 105, 327–332 (1999). https://doi.org/10.1007/s004399900135
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s004399900135