Abstract
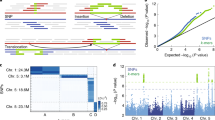
Single-nucleotide polymorphisms, as well as small insertions and deletions (here referred to collectively as simple nucleotide polymorphisms, or SNPs), comprise the largest set of sequence variants in most organisms1,2. Positional cloning based on SNPs may accelerate the identification of human disease traits and a range of biologically informative mutations3,4,5,6. The recent application of high-density oligonucleotide arrays to allele identification has made it feasible to genotype thousands of biallelic SNPs in a single experiment3,7. It has yet to be established, however, whether SNP detection using oligonucleotide arrays can be used to accelerate the mapping of traits in diploid genomes. The cruciferous weed Arabidopsis thaliana is an attractive model system for the construction and use of biallelic SNP maps. Although important biological processes ranging from fertilization and cell fate determination8,9,10,11 to disease resistance12,13 have been modelled in A. thaliana, identifying mutations in this organism has been impeded by the lack of a high-density genetic map consisting of easily genotyped DNA markers14. We report here the construction of a biallelic genetic map in A. thaliana with a resolution of 3.5 cM and its use in mapping Eds16, a gene involved in the defence response to the fungal pathogen Erysiphe orontii. Mapping of this trait involved the high-throughput generation of meiotic maps of F2 individuals using high-density oligonucleotide probe array-based genotyping. We developed a software package called InterMap and used it to automatically delimit Eds16 to a 7-cM interval on chromosome 1. These results are the first demonstration of biallelic mapping in diploid genomes and establish means for generalizing SNP-based maps to virtually any genetic organism.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Kruglyak, L. The use of a genetic map of biallelic markers in linkage studies. Nature Genet. 17, 21–24 (1997).
Kwok, P.Y., Deng, Q., Zakeri, H., Taylor, S.L. & Nickerson, D.A. Increasing the information content of STS-based genome maps: identifying polymorphisms in mapped STSs. Genomics 31, 123–126 (1996).
Wang, D.G. et al. Large-scale identification, mapping, and genotyping of single-nucleotide polymorphisms in the human genome. Science 280, 1077–1082 (1998).
Winzeler, E.A. et al. Direct allelic variation scanning of the yeast genome. Science 281, 1194–1197 ( 1998).
Hacia, J.G., Brody, L.C., Chee, M.S., Fodor, S.P. & Collins, F.S. Detection of heterozygous mutations in BRCA1 using high density oligonucleotide arrays and two-colour fluorescence analysis. Nature Genet. 14, 441– 447 (1996).
Buetow, K.H., Edmonson, M.N. & Cassidy, A.B. Reliable identification of large numbers of candidate SNPs from public EST data. Nature Genet. 21, 323–325 (1999).
Winzeler, E.A. & Davis, R.W. Functional analysis of the yeast genome. Curr. Opin. Genet. Dev. 7, 771– 776 (1997).
Sawa, S., Ito, T., Shimura, Y. & Okada, K. FILAMENTOUS FLOWER controls the formation and development of Arabidopsis inflorescences and floral meristems. Plant Cell 11, 69– 86 (1999).
Preuss, D., Lemieux, B., Yen, G. & Davis, R.W. A conditional sterile mutation eliminates surface components from Arabidopsis pollen and disrupts cell signaling during fertilization. Genes Dev. 7, 974–985 (1993).
Meyerowitz, E.M. Pattern formation in plant development: four vignettes. Curr. Opin. Genet. Dev. 4, 602–608 ( 1994).
Lloyd, A.M., Schena, M., Walbot, V. & Davis, R.W. Epidermal cell fate determination in Arabidopsis: patterns defined by a steroid-inducible regulator. Science 266, 436– 439 (1994).
Mindrinos, M., Katagiri, F., Yu, G.L. & Ausubel, F.M. The A. thaliana disease resistance gene RPS2 encodes a protein containing a nucleotide-binding site and leucine-rich repeats. Cell 78, 1089–1099 (1994).
Bent, A.F. et al. RPS2 of Arabidopsis thaliana: a leucine-rich repeat class of plant disease resistance genes. Science 265, 1856–1860 (1994).
Alonso-Blanco, C. et al. Development of an AFLP based linkage map of Ler, Col and Cvi Arabidopsis thaliana ecotypes and construction of a Ler/Cvi recombinant inbred line population. Plant J. 14, 259 –271 (1998).
Meinke, D.W., Cherry, J.M., Dean, C., Rounsley, S.D. & Koornneef, M. Arabidopsis thaliana: a model plant for genome analysis. Science 282, 662, 679– 682 (1998).
Hoogendoorn, B. et al. Genotyping single nucleotide polymorphisms by primer extension and high performance liquid chromatography. Hum. Genet. 104, 89–93 (1999).
O'Donovan, M.C. et al. Blind analysis of denaturing high-performance liquid chromatography as a tool for mutation detection. Genomics 52, 44–49 (1998).
Cronin, M.T. et al. Cystic fibrosis mutation detection by hybridization to light-generated DNA probe arrays. Hum. Mutat. 7, 244– 255 (1996).
Liu, Y.G., Mitsukawa, N., Lister, C., Dean, C. & Whittier, R.F. Isolation and mapping of a new set of 129 RFLP markers in Arabidopsis thaliana using recombinant inbred lines. Plant J. 10, 733–736 (1996).
Lander, E.S. et al. MAPMAKER: an interactive computer package for constructing primary genetic linkage maps of experimental and natural populations. Genomics 1, 174–181 ( 1987).
Konieczny, A. & Ausubel, F.M. A procedure for mapping Arabidopsis mutations using co-dominant ecotype-specific PCR-based markers. Plant J. 4, 403–410 ( 1993).
Li, J. & Chory, J. Preparation of DNA from Arabidopsis. Methods Mol. Biol. 82, 55– 60 (1998).
Lockhart, D.J. et al. Expression monitoring by hybridization to high-density oligonucleotide arrays. Nature Biotechnol. 14, 1675– 1680 (1996).
Chee, M. et al. Accessing genetic information with high-density DNA arrays. Science 274, 610–614 ( 1996).
Adam, L. & Somerville, S.C. Genetic characterization of five powdery mildew disease resistance loci in Arabidopsis thaliana. Plant J. 9, 341–356 ( 1996).
Volko, S.M., Boller, T. & Ausubel, F.M. Isolation of new Arabidopsis mutants with enhanced disease susceptibility to Pseudomonas syringae by direct screening. Genetics 149, 537–548 ( 1998).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Cho, R., Mindrinos, M., Richards, D. et al. Genome-wide mapping with biallelic markers in Arabidopsis thaliana . Nat Genet 23, 203–207 (1999). https://doi.org/10.1038/13833
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/13833
This article is cited by
-
Genome-Wide Identification and Expression Profiling of Tomato Invertase Genes Indicate Their Response to Stress and Phytohormones
Journal of Plant Growth Regulation (2022)
-
Identification and fine mapping of qSB.A09, a major QTL that controls shoot branching in Brassica rapa ssp. chinensis Makino
Theoretical and Applied Genetics (2020)
-
Fine mapping of an up-curling leaf locus (BnUC1) in Brassica napus
BMC Plant Biology (2019)
-
SNP genotyping in sheep from northwest and east China for meat traceability
Journal of Consumer Protection and Food Safety (2017)
-
Application of molecular markers in plant genome analysis: a review
The Nucleus (2017)