Summary
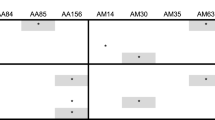
Gene flow by pollen between trees is essential for nut set in commercial almond orchards, due to the self-incompatibility of almond cultivars used. A study of gene flow has been carried out in an orchard composed of single rows of a “pollinating” cultivar between every double row of the most commercially desirable cultivar, Nonpareil. This “two-to-one” planting pattern was repeated throughout the orchard, and several “pollinating” cultivars were used in various parts of the orchard in an attempt to provide flowers for cross-pollination with Nonpareil at all stages of flowering of the latter. Using isozyme markers GPI-2, LAP-1, AAT-1, PGM-1, and PGM-2 and three newly-defined isozyme markers for almond — IDH, G6PD, and SDH — it has been shown that the gene flow resulting in nut set is quite restricted, taking place most strongly between neighboring halves of cross-compatibile pairs of trees. Even that half of a tree facing away from the “pollinating” tree has significantly less gene flow to it, while the next tree further on has few nuts set by fertilization from the “pollinating” tree under consideration. This result is surprising considering the comparatively large distances that the honeybee brought into the orchard in large numbers must travel within the orchard. To explain this apparent paradox and the observation that in most cases only a small proportion (<20%) of flowers set nuts, it is suggested that the honeybee predominantly visits only one cultivar, flying along the row of the cultivar to do so, and that cross-pollination results from accidental or rare visits involving two or more compatible cultivars.
Similar content being viewed by others
References
Baker BT, Gathercole F (1977) Commercial almond growing. South Australian Department of Agriculture Bulletin No. 9/77
Cerezo M, Socias i Company R, Arus P (1989) Identification of almond cultivars by pollen isozymes. J Am Soc Hortic Sci 114:164–169
Ellstrand NC, Derlin B, Marshall DL (1989) Gene flow by pollen into small populations: data from experimental and natural stands of wild radish. Proc Natl Acad Sci USA 86:9044–9047
Hauagge R, Kester DE, Asay RA (1987a) Isozyme variation among California almond cultivars. I. Inheritance. J Am Soc Hortic Sci 112:687–693
Hauagge R, Kester DE, Arulsekar S, Parfitt DE, Liu L (1987b) Isozyme variation among Californian almond cultivars. II. Cultivar characterization and origins. J Am Soc Hortic Sci 112:693–698
Hawker JS, Buttrose MS (1980) Development of the almond nut (Prunis dulcis (Mill) DA Webb). Anatomy and chemical composition of fruit parts from anthesis to maturity. Ann Bot 46:313–321
Hill SJ, Stephenson DW, Taylor BK (1985) Almond pollination studies: pollen production and viability, flower emergence and cross-pollination tests. Aust J Exp Agric 25:697–704
Jackson JF (1989) Borate control of protein secretion from Petunia pollen exhibits critical temperature discontinuities. Sex Plant Reprod 2:11–14
Kester DE, Griggs WH (1959) Fruit setting in the almond: the effect of cross-pollinating various percentages of flowers. Proc Am Soc Hortic Sci 74:206–213
Neale DB (1983) Dissertation Oregon State University, Corvallis/OR
Richardson BJ, Baverstock PR, Adams M (1986) Allozyme electrophoresis. A handbook for animal systematics and population studies. Academic Press Australia, Sydney, pp 99–144
Stanley RG, Linskens HF (1974) Pollen. Biology, biochemistry, management. Springer, Berlin Heidelberg New York, pp 87–115
Thorpe RW (1979) Honeybee foraging behavior in Californian almond orchards. In: Proc 4th Int Symp Pollination. Md Agric Exp Stn Misc Pub 1:385–390
Author information
Authors and Affiliations
Additional information
Communicated by H. F. Linskens
Rights and permissions
About this article
Cite this article
Jackson, J.F., Clarke, G.R. Gene flow in an almond orchard. Theoret. Appl. Genetics 82, 169–173 (1991). https://doi.org/10.1007/BF00226208
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/BF00226208