Summary
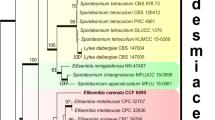
Mitochondrial genomes from yeasts in the Dekkera/Brettanomyces/Eeniella group vary in size from 28 to 101 kb. Mapping of genes has shown that the three smallest genomes, of 28–42 kb, have the same gene order, whereas the three larger mitochondrial DNAs of 57–101 kb are rearranged relative to the smaller molecules and between themselves. To examine the relationships between these genomes, a phylogenetic tree has been constructed by sequence comparison of the mitochondrialencoded cytochrome oxidase subunit gene (COX2) from the six species. Contrary to expectation, the tree shows that the larger rearranged genomes are more closely related than the smaller mtDNAs. This result indicates that the gene order of the smaller mtDNAs (28–42 kb) is ancestral and that larger mtDNA molecules (57–101 kb) are more prone to rearrangement than smaller forms.
Similar content being viewed by others
References
Brown WM, Prager EM, Wang A, Wilson AC (1982) Mitochondrial DNA sequences of primates: tempo and mode of evolution. J Mol Evol 18:225–239
Clark-Walker GD, McArthur CR, Sriprakash KS (1985) Location of transcriptional control signals and transfer RNA sequences in Torulopsis glabrata mitochondrial DNA. EMBO J 4:465–473
Clark-Walker GD, Hoeben P, Plazinska A, Smith DK, Wimmer E (1987) Application of mitochondrial DNA analysis to yeast systematics. In: De Hoog GS, Smith M Th, Weijman ACM (eds) The expanding realm of yeast-like fungi. Elsevier Science Publishers, Amsterdam, pp 259–266
Clark-Walker GD, McArthur CR, Daley DJ (1981) Does mitochondrial DNA length influence the frequency of spontaneous petite mutation in yeasts? Curr Genet 4:7–12
Clark-Walker GD (1989) In vivo rearrangement of mitochondrial DNA in Saccharomyces cerevisiae. Proc Natl Acad Sci USA 86:8847–8851
Clark-Walker GD (1992) Evolution of mitochondrial genomes in fungi. Int Rev Cytol (in press)
Curia R, Zalce ME, Mendoza V, Alvarez G, de Cobos AT, Brunner A (1990) Restriction site variation, length polymorphism and changes in gene order in the mitochondrial DNA of the yeast Kluyveromyces lactis. Ant Van Leeuwen 58:227–234
Coruzzi G, Tzagoloff A (1979) Assembly of the mitochondrial membrane system. DNA sequence of subunit 2 of yeast cytochrome oxidase. J Biol Chem 254:9324–9330
Coruzzi G, Bonitz SG, Thalenfeld BE, Tzagoloff A (1981) Assembly of the mitochondrial membrane system. Analysis of the nucleotide sequence and transcripts in the oxil region of yeast mitochondrial DNA. J Biol Chem 256:12780–12787
Feng DF, Johnson MS, Doolittle RF (1985) Aligning amino acid sequences: Comparison of commonly used methods. J Mol Evol 21:112–125
Feng DF, Doolittle RF (1987) Progressive sequence alignment as a prerequisite to correct phylogenetic trees. J Mol Evol 25: 351–360
Feng DF, Doolittle RF (1990) Progressive alignment and phylogenetic tree construction of protein sequences. Methods Enzymol 183:375–387
Fox TD (1987)Natural variation in the genetic code. Ann Rev Genet 21:67–91
Hardy CM, Clark-Walker GD (1990) Nucleotide sequence of the cytochrome oxidase subunit 2 and val-tRNA genes and surrounding sequences from Kluyveromyces lactis K8 mitochondrial DNA. Yeast 6:403–410
Hoeben P, Clark-Walker GD (1986) An approach to yeast classification by mapping mitochondrial DNA from Dekkeral/Brettanomyces and Eeniella genera. Curr Genet 10:371–379
Irwin DM, Kocher TD, Wilson AC (1991) Evolution of the cytochrome b gene of mammals. J Mol Evol 32:128–144
McArthur CR, Clark-Walker GD (1983) Mitochondrial DNA size diversity in the Dekkera/Brettanomyces yeasts. Curr Genet 7:29–35
Pratje E, Mannhaupt G, Michaelis G, Beyreuther K (1983) A nuclear mutation prevents processing of a mitochondrially encoded membrane protein in Saccharomyces cerevisiae. EMBO J 2:1049–1054
Saitou N, Nei M (1987) The neighbour joining method: A new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425
Sanger F, Nicklen S, Coulson AR (1977) DNA sequencing with chain termination inhibitors. Proc Natl Acad Sci USA 74: 5463–5467
Skelly PJ, Hardy CM, Clark-Walker GD (1991) A mobile group II intron of a naturally occurring rearranged mitochondrial genome in Kluyveromyces lactis. Curr Genet 20:115–120
Smith M Th, Batenburg-van der Vegte WH, Scheffers WA (1981) Eeniella, a new yeast genus of the Torulopsidales. Int J Syst Bacteriol 31:196–203
Smith M Th, Yamazaki M, Poot GA (1990) Dekkera, Brettanomyces and Eeniella: electrophoretic comparison of enzymes and DNA-DNA homology. Yeast 6:299–310
Studier JA, Keppler KJ (1988) A note on the neighbour-joining tree algorithm of Saitou and Nei. Mol Biol Evol 5:729–731
Author information
Authors and Affiliations
Additional information
Offprint requests to: G.D. Clark-Walker
Rights and permissions
About this article
Cite this article
Hoeben, P., Weiller, G. & Clark-Walker, G. Larger rearranged mitochondrial genomes in Dekkera/Brettanomyces yeasts are more closely related than smaller genomes with a conserved gene order. J Mol Evol 36, 263–269 (1993). https://doi.org/10.1007/BF00160482
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/BF00160482