Abstract
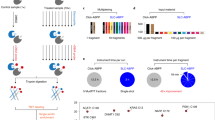
Proteomics research requires methods to characterize the expression and function of proteins in complex mixtures. Toward this end, chemical probes that incorporate known affinity labeling agents have facilitated the activity-based profiling of certain enzyme families. To accelerate the discovery of proteomics probes for enzyme classes lacking cognate affinity labels, we describe here a combinatorial strategy. Members of a probe library bearing a sulfonate ester chemotype were screened against complex proteomes for activity-dependent protein reactivity, resulting in the labeling of at least six mechanistically distinct enzyme classes. Surprisingly, none of these enzymes represented targets of previously described proteomics probes. The sulfonate library was used to identify an omega-class glutathione S-transferase whose activity was upregulated in invasive human breast cancer lines. These results indicate that activity-based probes compatible with whole-proteome analysis can be developed for numerous enzyme classes and applied to identify enzymes associated with discrete pathological states.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Anderson, N.L. & Anderson, N.G. Proteome and proteomics: new technologies, new concepts, and new words. Electrophoresis 19, 1853–1861 (1998).
Pandey, A. & Mann, M. Proteomics to study genes and genomes. Nature 405, 837–846 (2000).
Corthals, G.L., Wasinger, V.C., Hochstrasser, D.F. & Sanchez, J.C. The dynamic range of protein expression: a challenge for proteomic research. Electrophoresis 21, 1104–1115 (2000).
Nelson, P.S. et al. Comprehensive analyses of prostate gene expression: convergence of expressed sequence tag databases, transcript profiling and proteomics. Electrophoresis 21, 1823–1831 (2000).
Santoni, V., Molloy, M. & Rabilloud, T. Membrane proteins and proteomics: un amour impossible? Electrophoresis 21, 1054–1070 (2000).
Gygi, S.P., Corthals, G.L., Zhang, Y., Rochon, Y. & Aebersold, R. Evaluation of two-dimensional gel electrophoresis-based proteome analysis technology. Proc. Natl. Acad. Sci. USA 97, 9390–9395 (2000).
Legrain, P., Jestin, J.-L. & Schachter, V. From the analysis of protein complexes to proteome-wide linkage maps. Curr. Opin. Biotechnol. 11, 402–407 (2000).
Uetz, P. Two-hybrid arrays. Curr. Opin. Chem. Biol. 6, 57–62 (2002).
MacBeath, G. & Schreiber, S. Printing proteins as microarrays for high-throughput function determination. Science 289, 1760–1763 (2000).
Zhu, H. et al. Global analysis of protein activities using proteome chips. Science 293, 2101–2105 (2001).
Cravatt, B. & Sorensen, E. Chemical strategies for the global analysis of protein function. Curr. Opin. Chem. Biol. 4, 663–668 (2000).
Kidd, D., Liu, Y. & Cravatt, B.F. Profiling serine hydrolase activities in complex proteomes. Biochemistry 40, 4005–4015 (2001).
Greenbaum, D., Medzihradsky, K.F., Burlingame, A. & Bogyo, M. Epoxide electrophiles as activity-dependent cysteine protease profiling and discovery tools. Chem. Biol. 7, 569–581 (2000).
Liu, Y., Patricelli, M.P. & Cravatt, B.F. Activity-based protein profiling: the serine hydrolases. Proc. Natl. Acad. Sci. USA 96, 14694–14699 (1999).
Faleiro, L., Kobayashi, R., Fearnhead, H. & Lazebnik, Y. Multiple species of CPP32 and Mch2 are the major active caspases present in apoptotic cells. EMBO J. 16, 2271–2281 (1997).
Adam, G.C., Cravatt, B.F. & Sorensen, E.J. Profiling the specific reactivity of the proteome with non-directed activity-based probes. Chem. Biol. 8, 81–95 (2001).
Patricelli, M.P., Giang, D.K., Stamp, L.M. & Burbaum, J.J. Direct visualization of serine hydrolase activities in complex proteomes using fluorescent active site-directed probes. Proteomics 1, 1067–1071 (2001).
Penzes, P., Wang, X. & Napoli, J.L. Enzymatic characteristics of retinal dehydrogenase type I expressed in E. coli. Biochim. Biophys. Acta 1342, 175–181 (1997).
Hara, A. et al. Distribution and characterization of dihydrodiol dehydrogenases in mammalian ocular tissues. Biochem. J. 275, 113–119 (1991).
Rochefort, H. et al. Estrogen receptor mediated inhibition of cancer cell invasion and motility: an overview. J. Steroid Biochem. Mol. Biol. 65, 163–168 (1998).
Kodym, R., Calkins, P. & Story, M. The cloning and characterization of a new stress response protein: a mammalian member of a family of θ class glutathione-S-transferase-like proteins. J. Biol. Chem. 274, 5131–5137 (1999).
Hempel, J. et al. Aldehyde dehydrogenase catalytic mechanism: a proposal. Adv. Exp. Med. Biol. 7, 53–59 (1999).
Thompson, S. et al. Mechanistic studies on β-ketoacyl thiolase from Zoogloea ramigera: identification of the active-site nucleophile as Cys89, its mutation to Ser89, and kinetic and thermodynamic characterization of wild-type and mutant enzymes. Biochemistry 28, 5735–5742 (1989).
Board, P.G. et al. Identification, characterization, and crystal structure of the omega class glutathione transferases. J. Biol. Chem. 275, 24798–24806 (2000).
Armstrong, R.N. Kinetic and chemical mechanism of epoxide hydrolase. Drug Metab. Rev. 31, 71–86 (1999).
Terada, T. et al. Mutational analyses of cysteine residues of bovine dihydrodiol dehydrogenase 3. Biochim. Biophys. Acta 1547, 127–134 (2001).
Dakoji, S., Li, D., Agnihotri, G., Zhou, H.-Q. & Liu, H.-W. Studies on the inactivation of bovine liver enoyl-CoA hydratase by (methylenecyclopropyl)formyl-CoA: elucidation of the inactivation mechanism and identification of cysteine-114 as the entrapped nucleophile. J. Am. Chem. Soc. 123, 9749–9759 (2001).
Barrett, A.J. et al. L-trans-Epoxysuccinyl-leucylamido-(4-guanidino)butane (E-64) and its analogues as inhibitors of cysteine proteinases including cathepsins B, H and L. Biochem. J. 201, 189–198 (1982).
Dawson, J. & Holmes, C. Molecular mechanisms underlying inhibition of protein phosphatases by marine toxins. Front. Biosci. 4, D646–D658 (1999).
Acknowledgements
We thank G. Hawkins and M. Humphrey for technical assistance; J. Wu for assistance with mass spectrometry analysis; and J. Williamson, J. Kelly, and the Cravatt and Sorensen groups for helpful discussions. This work was supported by the National Cancer Institute of the National Institutes of Health (CA87660), the California Breast Cancer Research Program, ActivX Biosciences, and the Skaggs Institute for Chemical Biology.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
B.F.C. is a founder of Activx Biosciences, Inc., which has the right to license patents held by the Scripps Research Institute based on the technology described in this paper.
Rights and permissions
About this article
Cite this article
Adam, G., Sorensen, E. & Cravatt, B. Proteomic profiling of mechanistically distinct enzyme classes using a common chemotype. Nat Biotechnol 20, 805–809 (2002). https://doi.org/10.1038/nbt714
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt714
This article is cited by
-
An activity-dependent proximity ligation platform for spatially resolved quantification of active enzymes in single cells
Nature Communications (2017)
-
Mechanistic evaluation and transcriptional signature of a glutathione S-transferase omega 1 inhibitor
Nature Communications (2016)
-
Structure, function and disease relevance of Omega-class glutathione transferases
Archives of Toxicology (2016)
-
Chemical proteomic identification of T-plastin as a novel host cell response factor inHCV infection
Scientific Reports (2015)
-
Affinity purification in target identification: the specificity challenge
Archives of Pharmacal Research (2015)