Abstract
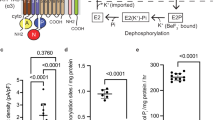
THE sodium pump or (Na+, K+)ATPase couples hydrolysis of ATP to the active transport of Na+ and K+ across the cell membrane. The firm association of the enzyme with the membrane allows its purification from the outer renal medulla by selective extraction of other membrane proteins using sodium dodecyl sulphate (SDS)1. Two polypeptide components remain in the purified preparation of (Na+, K+)ATPase. A catalytic protein with molecular weight (MW) about 100,000 (100 K) which constitutes 60–70% of the total protein and a smaller sialoglycoprotein of unknown function2,3. The arrangement of these proteins within the membrane is an important but poorly understood aspect of the Na pump structure. Presumably the catalytic regions of the enzyme are in contact with the aqueous phase at the membrane surface, while other portions of the protein are in intimate contact with the lipid and account for its firm anchorage. The lipid associated domains of the pump are of special interest because in addition to a purely structural role, they may also form parts of the ion conducting pathways. We have been able to identify presumptive intra-membranous portions of the Na pump protein by labelling them covalently from within the membrane with 5-[125I]-iodo-naphthyl-1-azide (INA). This label is a very hydrophobic light-sensitive compound developed previously for this purpose and tested with other systems4–7. A segment(s) of approximate MW 12K derived from the large polypeptide chain has been identified by combining the INA-labelling technique with extensive proteolysis of the membrane bound enzyme. Graded proteolysis of the labelled (Na+, K+)ATPase has also permitted an estimate of the position of this segment within the large chain.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Jorgensen, P. L. Biochim. biophys. Acta 356, 36–52 (1974).
Kyte, J. J. Biol. Chem. 247, 7642 (1972).
Jorgensen, P. L. Biochim. biophys. Acta 356, 53–67 (1974).
Klip, A. & Gitler, C. Biochem. biophys. Res. Commun. 60, 1155–1162 (1974).
Gitler, C. and Klip, A. in Perspectives in Membrane Biology (eds Estrado, O. S. & Gitler, C.) 149 (Academic, New York, 1974).
Sigrist-Nelson, K., Sigrist, H., Bercovici, T. & Gitler, C. Biochim. biophys. Acta 468, 163–176 (1977).
Bercovici, T. & Gitler, C. Biochemistry (submitted for publication).
Jorgensen, P. L. Biochim. biophys. Acta 401, 399–415 (1975).
Jorgensen, P. L. Biochim. biophys. Acta 446, 97–108 (1977).
Jorgensen, P. L. Meth. Enzym. 32 B, 277–290 (1974).
Karlish, S. J. D., Yates, D. W. & Glynn, I. M. Nature 263, 251–253 (1976).
Jensen, J. & Ottolenghi, P. Biochem. J. 159, 815–817 (1976).
Hegevary, C. & Post, R. L. J. biol. Chem. 246, 5234–5240 (1971).
Norby, J. G. & Jensen, J. Biochim. biophys. Acta 233, 104–116 (1971).
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
KARLISH, S., JORGENSEN, P. & GITLER, C. Identification of a membrane-embedded segment of the large polypeptide chain of (Na+, K+)ATPase. Nature 269, 715–717 (1977). https://doi.org/10.1038/269715a0
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/269715a0
This article is cited by
-
Structural basis for E1–E2 conformational transitions in Na, K-pump and Ca-pump proteins
The Journal of Membrane Biology (1988)
-
Localization of E1–E2 conformational transitions of sarcoplasmic reticulum Ca-ATPase by tryptic cleavage and hydrophobic labeling
The Journal of Membrane Biology (1986)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.