Abstract
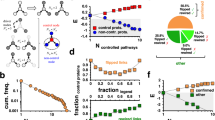
Two fundamental principles can account for how regulated networks of interacting proteins originated in cells. These are the law of mass action, which holds that the binding of one molecule to another increases with concentration, and the fact that the colocalization of molecules vastly increases their local concentrations. It follows that colocalization can amplify the effect on one protein of random mutations in another protein and can therefore, through natural selection, lead to interactions between proteins and to a startling variety of complex allosteric controls. It also follows that allostery is common and that homologous proteins can have different allosteric mechanisms. Thus, the regulated protein networks of organisms seem to be the inevitable consequence of natural selection operating under physical laws.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Hartwell, L. H., Hopfield, J. J., Leibler, S. & Murray, A. W. From molecular to modular cell biology. Nature 402, C47–C52 (1999).
Monod, J., Changeux, J. P. & Jacob, F. Allosteric proteins and cellular control systems. J. Mol. Biol. 6, 306–329 (1963).
Beernink, P. T., Endrizzi, J. A., Alber, T. & Schachman, H. K. Assessment of the allosteric mechanism of aspartate transcarbamoylase based on the crystalline structure of the unregulated catalytic subunit. Proc. Natl Acad. Sci. USA 96, 5388–5393 (1999).
Behe, M. J. Darwin's Black Box: The Biochemical Challenge to Evolution (The Free Press, New York, 2003).
Perutz, M. F. Stereochemistry of cooperative effects in haemoglobin. Nature 228, 726–739 (1970).
Xu, D., Tsai, C. J. & Nussinov, R. Mechanism and evolution of protein dimerization. Protein Sci. 7, 533–544 (1998).
Ispolatov, I., Yuryev, A., Mazo, I. & Maslov, S. Binding properties and evolution of homodimers in protein–protein interaction networks. Nucleic Acids Res. 33, 3629–3635 (2005).
Liang, J., Kim, J. R., Boock, J. T., Mansell, T. J. & Ostermeier, M. Ligand binding and allostery can emerge simultaneously. Protein Sci. 16, 929–937 (2007).
Pawson, T. & Scott, J. D. Signaling through scaffold, anchoring, and adaptor proteins. Science 278, 2075–2080 (1997).
Klemm, J. D. & Pabo, C. O. Oct-1 POU domain–DNA interactions: cooperative binding of isolated subdomains and effects of covalent linkage. Genes Dev. 10, 27–36 (1996).
Robinson, C. R. & Sauer, R. T. Covalent attachment of Arc repressor subunits by a peptide linker enhances affinity for operator DNA. Biochemistry 35, 109–116 (1996).
Predki, P. F. & Regan, L. Redesigning the topology of a four-helix-bundle protein: monomeric Rop. Biochemistry 34, 9834–9839 (1995).
Liang, H., Sandberg, W. S. & Terwilliger, T. C. Genetic fusion of subunits of a dimeric protein substantially enhances its stability and rate of folding. Proc. Natl Acad. Sci. USA 90, 7010–7014 (1993).
Pedersen, S., Bloch, P. L., Reeh, S. & Neidhardt, F. C. Patterns of protein synthesis in E. coli: a catalog of the amount of 140 individual proteins at different growth rates. Cell 14, 179–190 (1978).
Lu, P., Vogel, C., Wang, R., Yao, X. & Marcotte, E. M. Absolute protein expression profiling estimates the relative contributions of transcriptional and translational regulation. Nature Biotechnol. 25, 117–124 (2007).
Noguchi, C. T. & Schechter, A. N. Sickle hemoglobin polymerization in solution and in cells. Annu. Rev. Biophys. Biophys. Chem. 14, 239–263 (1985).
Fermi, G., Perutz, M. F., Shaanan, B. & Fourme, R. The crystal structure of human deoxyhaemoglobin at 1.74 Å resolution. J. Mol. Biol. 175, 159–174 (1984).
Bahadur, R. P., Chakrabarti, P., Rodier, F. & Janin, J. Dissecting subunit interfaces in homodimeric proteins. Proteins 53, 708–719 (2003).
Fersht, A. R. Structure and Mechanism in Protein Science (Freeman, New York, 1999).
Eisenberg, D., Wesson, M. & Yanashita, M. Interpretation of protein folding and binding with atomic solvation parameters. Chem. Scr. 29A, 217–221 (1989).
Behe, M. J. The Edge of Evolution (Free Press, New York, 2007).
Marcotte, E. M., Pellegrini, M., Thompson, M. J., Yeates, T. O. & Eisenberg, D. A combined algorithm for genome-wide prediction of protein function. Nature 402, 83–86 (1999).
Bennett, M. J., Choe, S. & Eisenberg, D. Domain swapping: entangling alliances between proteins. Proc. Natl Acad. Sci. USA 91, 3127–3131 (1994).
Schlunegger, M. P., Bennett, M. J. & Eisenberg, D. Oligomer formation by 3D domain swapping: a model for protein assembly and misassembly. Adv. Protein Chem. 50, 61–122 (1997).
Finn, R. D. et al. Pfam: clans, web tools and services. Nucleic Acids Res. 34, D247–D251 (2006).
Servant, F. et al. ProDom: automated clustering of homologous domains. Brief. Bioinform. 3, 246–251 (2002).
Enright, A. J., Iliopoulos, I., Kyrpides, N. C. & Ouzounis, C. A. Protein interaction maps for complete genomes based on gene fusion events. Nature 402, 86–90 (1999).
Thoden, J. B., Raushel, F. M., Benning, M. M., Rayment, I. & Holden, H. M. The structure of carbamoyl phosphate synthetase determined to 2.1 Å resolution. Acta Crystallogr. D Biol. Crystallogr. 55, 8–24 (1999).
Snel, B., Bork, P. & Huynen, M. Genome evolution: gene fusion versus gene fission. Trends Genet. 16, 9–11 (2000).
Kummerfeld, S. K. & Teichmann, S. A. Relative rates of gene fusion and fission in multi-domain proteins. Trends Genet. 21, 25–30 (2005).
Fong, J. H., Geer, L. Y., Panchenko, A. R. & Bryant, S. H. Modeling the evolution of protein domain architectures using maximum parsimony. J. Mol. Biol. 366, 307–315 (2007).
Pabo, C. O. & Sauer, R. T. Transcription factors: structural families and principles of DNA recognition. Annu. Rev. Biochem. 61, 1053–1095 (1992).
Wolberger, C. Multiprotein–DNA complexes in transcriptional regulation. Annu. Rev. Biophys. Biomol. Struct. 28, 29–56 (1999).
Panne, D., Maniatis, T. & Harrison, S. C. An atomic model of the interferon-β enhanceosome. Cell 129, 1111–1123 (2007).
Wilson, D. S., Guenther, B., Desplan, C. & Kuriyan, J. High resolution crystal structure of a paired (Pax) class cooperative homeodomain dimer on DNA. Cell 82, 709–719 (1995).
Passner, J. M., Ryoo, H. D., Shen, L., Mann, R. S. & Aggarwal, A. K. Structure of a DNA-bound Ultrabithorax–Extradenticle homeodomain complex. Nature 397, 714–719 (1999).
Piper, D. E., Batchelor, A. H., Chang, C. P., Cleary, M. L. & Wolberger, C. Structure of a HoxB1–Pbx1 heterodimer bound to DNA: role of the hexapeptide and a fourth homeodomain helix in complex formation. Cell 96, 587–597 (1999).
LaRonde-LeBlanc, N. A. & Wolberger, C. Structure of HoxA9 and Pbx1 bound to DNA: Hox hexapeptide and DNA recognition anterior to posterior. Genes Dev. 17, 2060–2072 (2003).
Klemm, J. D., Rould, M. A., Aurora, R., Herr, W. & Pabo, C. O. Crystal structure of the Oct-1 POU domain bound to an octamer site: DNA recognition with tethered DNA-binding modules. Cell 77, 21–32 (1994).
Jacobson, E. M., Li, P., Leon- del-Rio, A., Rosenfeld, M. G. & Aggarwal, A. K. Structure of Pit-1 POU domain bound to DNA as a dimer: unexpected arrangement and flexibility. Genes Dev. 11, 198–212 (1997).
Li, T., Stark, M. R., Johnson, A. D. & Wolberger, C. Crystal structure of the Mata1/MATα2 homeodomain heterodimer bound to DNA. Science 270, 262–269 (1995).
Tan, S. & Richmond, T. J. Crystal structure of the yeast MATα2/MCM1/DNA ternary complex. Nature 391, 660–666 (1998).
Royer, W. E., Zhu, H., Gorr, T. A., Flores, J. F. & Knapp, J. E. Allosteric hemoglobin assembly: diversity and similarity. J. Biol. Chem. 280, 27477–27480 (2005).
Royer, W. E., Hendrickson, W. A. & Chiancone, E. The 2.4 Å crystal structure of Scapharca dimeric hemoglobin. J. Biol. Chem. 264, 21052–21061 (1989).
Barford, D., Hu, S. H. & Johnson, L. N. Structural mechanism for glycogen phosphorylase control by phosphorylation and AMP. J. Mol. Biol. 218, 233–260 (1991).
Barford, D. & Johnson, L. N. The allosteric transition of glycogen phosphorylase. Nature 340, 609–616 (1989).
Sprang, S. R. et al. Structural changes in glycogen phosphorylase induced by phosphorylation. Nature 336, 215–221 (1988).
Buchbinder, J. L., Rath, V. L. & Fletterick, R. J. Structural relationships among regulated and unregulated phosphorylases. Annu. Rev. Biophys. Biomol. Struct. 30, 191–209 (2001).
Rath, V. L. & Fletterick, R. J. Parallel evolution in two homologues of phosphorylase. Nature Struct. Biol. 1, 681–690 (1994).
Palm, D., Goerl, R. & Burger, K. J. Evolution of catalytic and regulatory sites in phosphorylases. Nature 313, 500–502 (1985).
Hwang, P. K. & Fletterick, R. J. Convergent and divergent evolution of regulatory sites in eukaryotic phosphorylases. Nature 324, 80–84 (1986).
Lee, S. Y. et al. Regulation of the transcriptional activator NtrC1: structural studies of the regulatory and AAA+ ATPase domains. Genes Dev. 17, 2552–2563 (2003).
Doucleff, M. et al. Negative regulation of AAA+ ATPase assembly by two component receiver domains: a transcription activation mechanism that is conserved in mesophilic and extremely hyperthermophilic bacteria. J. Mol. Biol. 353, 242–255 (2005).
De Carlo, S. et al. The structural basis for regulated assembly and function of the transcriptional activator NtrC. Genes Dev. 20, 1485–1495 (2006).
Pawson, T. & Nash, P. Assembly of cell regulatory systems through protein interaction domains. Science 300, 445–452 (2003).
Neet, K. & Hunter, T. Vertebrate non-receptor protein-tyrosine kinase families. Genes Cells 1, 147–169 (1996).
Nagar, B. et al. Structural basis for the autoinhibition of c-Abl tyrosine kinase. Cell 112, 859–871 (2003).
Sicheri, F., Moarefi, I. & Kuriyan, J. Crystal structure of the Src-family tyrosine kinase Hck. Nature 385, 602–609 (1997).
Xu, W., Harrison, S. C. & Eck, M. J. Three-dimensional structure of the tyrosine kinase c-Src. Nature 385, 595–602 (1997).
Brasher, B. B. & Van Etten, R. A. c-Abl has high intrinsic tyrosine kinase activity that is stimulated by mutation of the Src homology 3 domain and by autophosphorylation at two distinct regulatory sites. J. Biol. Chem. 275, 35631–35637 (2000).
Deindl, S. et al. Structural basis for the inhibition of tyrosine kinase activity of ZAP-70. Cell 129, 735–746 (2007).
Lietha, D. et al. Structural basis for the autoinhibition of focal adhesion kinase. Cell 129, 1177–1187 (2007).
Schlessinger, J. Cell signaling by receptor tyrosine kinases. Cell 103, 211–225 (2000).
Schlessinger, J. Ligand-induced, receptor-mediated dimerization and activation of EGF receptor. Cell 110, 669–672 (2002).
Hubbard, S. R. & Till, J. H. Protein tyrosine kinase structure and function. Annu. Rev. Biochem. 69, 373–398 (2000).
Zhang, X., Gureasko, J., Shen, K., Cole, P. A. & Kuriyan, J. An allosteric mechanism for activation of the kinase domain of epidermal growth factor receptor. Cell 125, 1137–1149 (2006).
Deeds, E. J., Ashenberg, O., Gerardin, J. & Shakhnovich, E. I. Robust protein–protein interactions in crowded cellular environments. Proc. Natl Acad. Sci. USA 104, 14952–14957 (2007).
Royer, W. E. . High-resolution crystallographic analysis of a co-operative dimeric hemoglobin. J. Mol. Biol. 235, 657–681 (1994).
Flores, J. F. et al. Sulfide binding is mediated by zinc ions discovered in the crystal structure of a hydrothermal vent tubeworm hemoglobin. Proc. Natl Acad. Sci. USA 102, 2713–2718 (2005).
Lin, K., Rath, V. L., Dai, S. C., Fletterick, R. J. & Hwang, P. K. A protein phosphorylation switch at the conserved allosteric site in GP. Science 273, 1539–1542 (1996).
Cho, H. S. & Leahy, D. J. Structure of the extracellular region of HER3 reveals an interdomain tether. Science 297, 1330–1333 (2002).
Garrett, T. P. et al. Crystal structure of a truncated epidermal growth factor receptor extracellular domain bound to transforming growth factor α. Cell 110, 763–773 (2002).
Ogiso, H. et al. Crystal structure of the complex of human epidermal growth factor and receptor extracellular domains. Cell 110, 775–787 (2002).
Acknowledgements
We are grateful to A. K. Aggarwal, R. E. Dickerson, S. C. Harrison, L. N. Johnson, E. M. Marcotte, M. Robertson, W. E. Royer, M. Seeliger, D. E. Wemmer, C. Wolberger, T. O. Yeates and many other colleagues for comments. We thank S. Deindl, L. Leighton, W. E. Royer, D. E, Wemmer and X. Zhang for assistance with the figures. Support from the Howard Hughes Medical Institute, the National Institutes of Health, the US Department of Energy Office of Biological & Environmental Research, and the National Science Foundation is gratefully acknowledged.
Author information
Authors and Affiliations
Additional information
Correspondence should be addressed to the authors (david@mbi.ucla.edu; kuriyan@berkeley.edu).
Rights and permissions
About this article
Cite this article
Kuriyan, J., Eisenberg, D. The origin of protein interactions and allostery in colocalization. Nature 450, 983–990 (2007). https://doi.org/10.1038/nature06524
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature06524
This article is cited by
-
Unified mRNA Subcellular Localization Predictor based on machine learning techniques
BMC Genomics (2024)
-
MSLP: mRNA subcellular localization predictor based on machine learning techniques
BMC Bioinformatics (2023)
-
Insight into de-regulation of amino acid feedback inhibition: a focus on structure analysis method
Microbial Cell Factories (2023)
-
Functional advantages of building nanosystems using multiple molecular components
Nature Chemistry (2023)
-
Impact of oligomerization on the allergenicity of allergens
Clinical and Molecular Allergy (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.