Summary
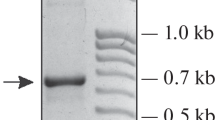
A complete ribosomal DNA (rDNA) repeat unit has been cloned from the genome of Pisum sativum (garden pea) and used to construct a map containing a total of 58 cleavage sites for 23 different restriction enzymes. Regions encoding 18s and 25s ribosomal RNA (rRNA) were identified by R-loop analysis. A 180 bp sequence element is repeated eight times in the intergenic ‘nontranscribed spacer’ (NTS) region, as defined by eight evenly spaced RsaI cleavage sites. Sequence heterogeneity among these elements (subrepeats) is indicated by the presence of an NcoI site within the five RsaI subrepeats distal to the 25s rRNA gene but not in the three subrepeats proximal to this gene, and also by the presence of an additional RsaI cleavage site in one subrepeat.
The approximately 4000 copies of the rDNA repeat in the pea nuclear genome show considerable heterogeneity with respect to the length of the NTS region, and differences are also frequently observed between different genotypes. In both cases the length variation appears to be due primarily to differences in the number of subrepeat elements.
Comparison of rDNA restriction maps for two pea genotypes separated for hundreds or perhaps thousands of generations reveals that they contain many rDNA identical repeat units. This data is consistent with the view that new rDNA variants are fixed only infrequently in the evolution of a species.
Differences also exist between the rDNA repeats of a single genotype with respect to the degree of base modification at certain restriction sites. A large number of sites known to exist in the pea rDNA clone are not cleaved at all in genomic rDNA, or are cleaved in only some copies of the rDNA repeat. We believe these examples of incomplete cleavage results mostly from methylation, although it is difficult to rule out the possibility of sequence variation in all cases. Most putative modifications are best interpreted in terms of cytosine methylation in CG and CXG sequences, but at least one example is more consistent with adenine methylation.
We also have constructed a more detailed restriction map of the wheat rDNA clone pTA71 and present a comparison of this map to our map of pea, pumpkin, and wheat in order to assess the amount of useful evolutionary information that can be obtained by comparison of such maps.
Similar content being viewed by others
References
Appels R, Dvorak J: Relative rates of divergence of spacer and gene sequences within the rDNA region of species in the Triticeae: Implications for the maintenance of homogeneity of a repeated gene family. Theor Appl Genet 63:361–365, 1982.
Chang AC, Cohen SN: Construction and characterization of amplifiable multicopy DNA cloning vehicles derived from the P15A cryptic miniplasmid. Journal of Bacteriology 143:1141–1156, 1978.
Coen ES, Dover GA: Unequal exchanges and the coevolution of X and Y rDNA arrays in Drosophila melanogaster. Cell 33:849–855, 1983.
Coen ES, Thoday JM, Dover G: Rate of turnover of structural variants in the rDNA gene family of Drosophila melanogaster. Nature 295:564–567, 1982.
Dover GA, Flavell RB: Molecular coevolution: DNA diver gence and the maintenance of function. Cell 38:622–623, 1984.
Dover GA, Brown SDM, Coen ES, Dallas J, Strachan T, Trick M: The dynamics of genome evolution and species differentiation. In: Dover GA, Flavell RB (eds) Genome Evolution. Academic Press, London, 1982, pp 343–372.
Eckenrode VK, Arnold J, Meagher RB: Comparison of the sequence of soybean 18S rRNA with the sequences of other small subunit rRNAs. J Mol Evol 21:259–269, 1985.
Ellis THN, Davies DR, Castleton JA, Bedford ID: The organization and genetics of rDNA length variants in peas. Chromosoma 91:74–81, 1984.
Flavell RB: The structure and control of expression of ribosomal RNA genes. In: Miflin B (ed) Oxford Surveys of Plant Molecular and Cell Biology, Volume 3. Oxford University Press, Oxford, England (in press) 1986.
Flavell RB, O'Dell M, Thompson WF: Cytosine methylation of ribosomal RNA genes and nucleolus organizer activity in wheat. Proc Kew Chromosome Conf II:11–17, 1983.
Flavell RB, O'Dell M, Smith DB, Thompson WF: Chromosome architecture: the distribution of recombination sites, the structure of ribosomal DNA loci, and the multiplicity of sequences containing inverted repeats. In: vanVloten-Doting L, Groot GSP, Hall TC (eds) Molecular Form and Function of the Plant Genome. Plenum Press, New York, 1985.
Gerbi SA, Gourse RL, Clark CG: Conserved regions within ribosomal DNA: Locations and some possible functions. The Cell Nucleus. Academic Press, Inc., 1982.
Gerlach WL, Bedbrook JR: Cloning and characterization of ribosomal RNA genes from wheat and barley. Nucleic Acids Res 7:1869–1879, 1979.
Gruenbaum Y, Naveh-Many T, Cedar H, Razin A: Sequence specificity of methylation in higher plant DNA. Nature 292:860–862, 1981.
Grunstein M, Hogness DS: Colony hybridization: A method for the isolation of cloned DNAs that contain a specific gene. Proc Nat Acad Sci USA 72:3961–3965, 1975.
Ingle I, Timmis IN, Sinclair J: The relationship between satellite desoxyribonucleic acid, ribosomal ribonucleic acid gene redundancy and genome size in plants. Plant Physiol 55: 496–501, 1975.
Jorgensen RA, Cuellar RE, Thompson WF: Modes and tempos in the evolutions of nuclear-encoded ribosomal RNA genes in legumes. Carnegie Inst Wash Year Book 81:98–101, 1982.
Kohorn BD, Rae PMM: Nontranscribed spacer sequences promote in vitro transcription of Drosophila ribosomal DNA. Nucleic Acids Res 10:6879–6886, 1982.
Maniatis T, Fritsch EF, Sambrook J: Molecular Cloning. Cold Spring Harbor Laboratory, New York, 1982.
McClelland M: The effect of sequence specific DNA methylation on restriction endonuclease cleavage (update). Nucleic Acids Res 9:5859–5865, 1981.
McClelland M: The effect of site specific methylation on restriction endonuclease cleavage (update). Nucleic Acids Res 11:r169–172, 1983.
McClelland M, Nelson M: The effect of site specific methylation on restriction endonuclease digestion. Nucleic Acids Res 13:r201–207, 1985.
Messing J, Carlson J, Hagen G, Rubenstein I, Oleson A: Cloning and sequencing the ribosomal RNA genes in maize: the 17S region. DNA 3:31–40, 1984.
Moss T: A transcriptional function for the repetitive spacer in Xenopus laevis. Nature 302:223–230, 1983.
Murray MG, Thompson WF: Rapid isolation of high molecular weight plant DNA. Nucleic Acids Res 8:4231–4236, 1980.
Murtif VL, Rae PMM: In vivo transcription of rDNA spacers in Drosophila. Nucleic Acids Res 13:3221–3239, 1985.
Oono K, Sugiura M: Heterogeneity of the ribosomal RNA gene clusters in rice. Chromosoma 76:85–89, 1980.
Palmer JD, Jorgensen RA, Thompson WF: Chloroplast DNA variation and evolution in Pisum: Patterns of change and phylogenetic analysis. Genetics 108:195–213, 1985.
Polans NO, Weeden NF, Thompson WF: Distribution, inheritance and linkage relationships of ribosomal DNA spacer length variants in pea. Theor Appl Genet, in press.
Rafalski JA, Wiewjiorowski M, Soll D: Organization of ribosomal DNA in yellow lupine (Lupinus luteus) and sequence of the 5.8S RNA gene. FEBS Lett 241–244, 1983.
Reeder RH: Enhancers and ribosomal gene spacers. Cell 38:349–351, 1984.
Saghai-Maroof MA, Soliman KM, Jorgensen RA, Allard RW: Ribosomal DNA spacer-length polymorphisms in barley: Mendelian inheritance, chromosomal location, and population dynamics. Proc Natl Acad Sci USA 81:8014–8018, 1984.
Siegel A, Kolacz K: Heterogeneity of pumpkin ribosomal DNA. Plant Physiol 72:166–171, 1983.
Smith HO, Birnstiel ML: A simple method for DNA restriction site mapping. Nucleic Acids Res 3:2387–2395, 1976.
Treco D, Brownell E, Arnhe N: The ribosomal gene nontranscribed spacer. In: The Cell Nucleus, Academic Press, Inc., 1982.
Waldron J, Dunsmuir P, Bedbrook J: Characterization of the rDNA repeat units in the Mitchell Petunia genome. Plant Mol Biol 2:57–65, 1983.
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Jorgensen, R.A., Cuellar, R.E., Thompson, W.F. et al. Structure and variation in ribosomal RNA genes of pea. Plant Mol Biol 8, 3–12 (1987). https://doi.org/10.1007/BF00016429
Received:
Revised:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/BF00016429