Abstract
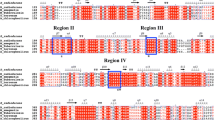
We cloned the Kluyveromyces lactis KlNTH1 gene, which encodes neutral trehalase. It showed 65.2% and 68.5% identity at nucleotide and amino acid sequence level, respectively, with the Saccharomyces cerevisiae NTH1 gene. Multiple alignment of the predicted trehalase protein sequences from yeasts, bacteria, insects, and mammals revealed two major domains of conservation. Only the yeast trehalases displayed in an N-terminal extension two consensus sites for cAMP-dependent protein phosphorylation and a putative Ca2+-binding sequence. Gene disruption of the KlNTH1 gene abolished neutral trehalase activity and clearly revealed a trehalase activity with an acid pH optimum. It also resulted in a high constitutive trehalose level. Expression of the KlNTH1 gene in an S. cerevisiae nth1Δ mutant resulted in rapid activation of the heterologous trehalase upon addition of glucose to cells growing on a nonfermentable carbon source and upon addition of a nitrogen source to cells starved for nitrogen in a glucose-containing medium. In K. lactis, the same responses were observed except that rapid activation by glucose was observed only in early-exponential-phase cells. Inactivation of K. lactis neutral trehalase by alkaline phosphatase and activation by cAMP in cell extracts are consistent with control of the enzyme by cAMP-dependent protein phosphorylation.
Similar content being viewed by others
Author information
Authors and Affiliations
Additional information
Received: 19 March 1996 / Accepted: 15 October 1996
Rights and permissions
About this article
Cite this article
Amaral, F., Dijck, P., Nicoli, J. et al. Molecular cloning of the neutral trehalase gene from Kluyveromyces lactis and the distinction between neutral and acid trehalases. Arch Microbiol 167, 202–208 (1997). https://doi.org/10.1007/s002030050436
Published:
Issue Date:
DOI: https://doi.org/10.1007/s002030050436



