Abstract
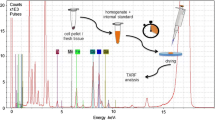
Total Reflection X-Ray Fluorescence Spectrometry (TXRF) offers many advantages for the detection of trace elements in enzymes as compared to other well known analytical techniques like flame-AAS or ICP-AES because of the significantly smaller amounts of sample required. Without any decomposition, elements like Fe, Ni, Cu, Zn, Mn and Mo could be determined with high accuracy, in spite of the large bio-organic matrix. Besides the metals also sulfur can be determined in protein samples. The two terminal oxidases, cytochrome c oxidase and quinol oxidase, isolated from the soil bacterium Paracoccus denitrificans, were transferred from their usual salt buffer into a solution of 100 mmol/L tris(hydroxymethyl)aminomethane (TRIS) acetate containing an appropriate detergent. By this procedure an improved signal/noise ratio is attained. The data for cytochrome c oxidase are in good agreement with values obtained by ICP-AES. Further results of quinol oxidase, which has different element ratios, also fit the expected values. The investigations lead to the conclusion that the method is well suited for the quantitative determination of metals in enzymes, and in particular their molar ratios, and requires only small amounts of the biological sample without any extensive pretreatment.
Similar content being viewed by others
Author information
Authors and Affiliations
Additional information
Received: 17 June 1997 / Revised: 21 November 1997 / Accepted: 27 November 1997
Rights and permissions
About this article
Cite this article
Wittershagen, A., Rostam-Khani, P., Zickermann, V. et al. Determination of metal-cofactors in respiratory chain complexes by total-reflection X-ray fluorescence spectrometry (TXRF). Fresenius J Anal Chem 361, 326–328 (1998). https://doi.org/10.1007/s002160050895
Issue Date:
DOI: https://doi.org/10.1007/s002160050895