Summary
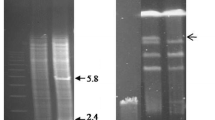
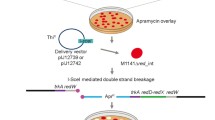
The genes SNQ and SFA confer hyperresistance to 4-NQO and FA when present on a multi-copy plasmid in yeast. Both are non-essential genes since transplacement of SNQ by a disrupted snq-0::LEU2 yielded stable and viable haploid integrants. Southern analysis revealed that SNQ and SFA are single-loci genes, and OFAGE analysis showed that they are located on chromosome XIII and IV, respectively. Northern blot analysis of SNQ and SFA revealed poly(A)+ RNA transcripts of 2 kb and 1.7 kb, respectively. Nuclease S 1 mapping showed SNQ to have a coding region of 1.6 kb and SFA, one of 1.3 kb. The 5′ coding regions were determined for both genes, while the 3′ end could only be determined for gene SNQ. Both genes do not appear to contain introns. The SFA locus was also mapped by transposon mutagenesis. Tn10-LUK integrants disrupted the SFA gene function at sites that were determined by subcloning to lie within the SFA transcription unit.
Similar content being viewed by others
Abbreviations
- 4-NQO :
-
4-Nitro-quinoline-N-oxide
- FA :
-
formaldehyde
References
Birnboim HG, Doly J (1979) Nucleic Acids Res 7:1513–1523
Clifton D, Weinstock SB, Fraenkel DG (1978) Genetics 88:1–11
Dagert M, Ehrlich SD (1979) Gene 6:23–28
Favoloro J, Treisman R, Kamen R (1980) Methods Enzymol 65:718–749
Fleer R, Brendel M (1979) Mol Gen Genet 176:41–52
Hinnen A, Hicks JB, Fink GR (1978) Proc Natl Acad Sci USA75:1919–1933
Holm C, Meeks-Wagner DW, Fangman WL, Botstein D (1986) Gene 42:169–173
Huisman O, Raymond W, Froehlich KU, Errada P, Kleckner N, Botstein D, Hoyt MA (1987) Genetics 116:191–199
Ikenaga M, Ichikawa-Ryo H, Kondo S (1975) J Mol Biol 92:341–356
Ito H, Fukuda J, Murata K, Kimura A (1983) J Bacteriol 153:163–168
Kondo S, Ichikawa H, Iwo K, Kato T (1970) Genetics 66:187–217
Mack M, Gömpel-Klein P, Haase E, Hietkamp J, Ruhland At, Brendel M (1988) Mol Gen Genet 211:260–265
Maniatis T, Fritsch EF, Sambrook J (1982) Molecular cloning — a laboratory manual. Cold Spring Harbor Laboratory, Cold Spring Harbor, NY
Miller JA, Miller EC (1976) In: Montesano R, Bartsch H, Tomatis L (eds) Screening tests in chemical carcinogenesis. Int Agency for Research on Cancer, Lyon, pp 153–180
Miller JH (1972) Experiments in molecular genetics. Cold Spring Harbor Laboratory, Cold Spring Harbor, NY
Nicolet CM, Chenevert JM, Friedberg EC (1985) Gene 36:225–234
Prakash L (1976) Genetics 83:285–301
Prakash L, Sherman F (1973) J Mol Biol 79:65–82
Rodriguez RL, Tait RC (1983) In: Recombinant DNA techniques (an introduction). Addison-Wesley Publ. Co, London, pp 170–172
Rothstein RJ (1983) Methods Enzymol 101:202–209
Ruhland AR, Brendel M (1979) Genetics 92:83–97
Ruhland AR, Haase E, Siede W, Brendel M (1981) Mol Gen Genet 181:346–351
Ruhland AR, Brendel M, Haynes RH (1986) Curr Genet 11:211–215
Sander P, Brendel M (1988) Curt Genet 12:125–128
Schultz LD, Friesen JD (1983) J Bacteriol 155:8–14
Seelmann K (1988) Physiologische Grundlagen der Formaldehyd-Hyperresistenz in Bäckerhefe. Diplomarbeit FB Biologie der J W. Goethe-Universität, Frankfurt/Main
Siede W (1986) Analyse mutagener DNA-Reparatur in Saccharomyces cerevisiae: Charakterisierung des REV2 abhängigen Prozesses. Dissertation FB Biologie der J. W. Goethe-Universität Frankfurt/Main
Sugimura T, Okabe K, Nagao M (1966) Cancer Res 26:1717–1721
Tada M (1981) Carcinogenesis 6:25–45
Tada M, Tada M (1975) Nature 255:510–512
Way JC, Davis MA, Mosisato D, Roberts DE, Kleckner N (1984) Gene 32:369–379
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Gömpel-Klein, P., Mack, M. & Brendel, M. Molecular characterization of the two genes SNQ and SFA that confer Hyperresistance to 4-nitroquinoline-N-oxide and formaldehyde in Saccharomyces cerevisiae . Curr Genet 16, 65–74 (1989). https://doi.org/10.1007/BF00393397
Received:
Issue Date:
DOI: https://doi.org/10.1007/BF00393397