Abstract
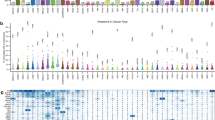
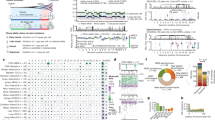
Cancers arise owing to mutations in a subset of genes that confer growth advantage. The availability of the human genome sequence led us to propose that systematic resequencing of cancer genomes for mutations would lead to the discovery of many additional cancer genes. Here we report more than 1,000 somatic mutations found in 274 megabases (Mb) of DNA corresponding to the coding exons of 518 protein kinase genes in 210 diverse human cancers. There was substantial variation in the number and pattern of mutations in individual cancers reflecting different exposures, DNA repair defects and cellular origins. Most somatic mutations are likely to be ‘passengers’ that do not contribute to oncogenesis. However, there was evidence for ‘driver’ mutations contributing to the development of the cancers studied in approximately 120 genes. Systematic sequencing of cancer genomes therefore reveals the evolutionary diversity of cancers and implicates a larger repertoire of cancer genes than previously anticipated.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Futreal, P. A. et al. A census of human cancer genes. Nature Rev. Cancer 4, 177–183 (2004)
Futreal, P. A. et al. Cancer and genomics. Nature 409, 850–852 (2001)
Sawyers, C. Targeted cancer therapy. Nature 432, 294–297 (2004)
Manning, G., Whyte, D. B., Martinez, R., Hunter, T. & Sudarsanam, S. The protein kinase complement of the human genome. Science 298, 1912–1934 (2002)
Stephens, P. et al. A screen of the complete protein kinase gene family identifies diverse patterns of somatic mutations in human breast cancer. Nature Genet. 37, 590–592 (2005)
Davies, H. et al. Somatic mutations of the protein kinase gene family in human lung cancer. Cancer Res. 65, 7591–7595 (2005)
Hunter, C. et al. A hypermutation phenotype and somatic MSH6 mutations in recurrent human malignant gliomas after alkylator chemotherapy. Cancer Res. 66, 3987–3991 (2006)
Bignell, G. et al. Sequence analysis of the protein kinase gene family in human testicular germ-cell tumours of adolescents and adults. Genes Chromosom. Cancer 45, 42–46 (2006)
Bardelli, A. et al. Mutational analysis of the tyrosine kinome in colorectal cancers. Science 300, 949 (2003)
Wang, T.-L. et al. Prevalence of somatic alterations in the colorectal cancer cell genome. Proc. Natl Acad. Sci. USA 99, 3076–3080 (2002)
Lonardi, S., Tosoni, A. & Brandes, A. A. Adjuvant chemotherapy in the treatment of high grade gliomas. Cancer Treat. Rev. 31, 79–89 (2005)
Karran, P., Offman, J. & Bignami, M. Human mismatch repair, drug-induced DNA damage, and secondary cancer. Biochimie 85, 1149–1160 (2003)
Greenman, C., Wooster, R., Futreal, P. A., Stratton, M. R. & Easton, D. F. Statistical analysis of pathogenicity of somatic mutations in cancer. Genetics 173, 2187–2198 (2006)
Yang, Z., Ro, S. & Rannala, B. Likelihood models of somatic mutation and codon substitution in cancer genes. Genetics 165, 695–705 (2003)
Goldman, N. & Yang, Z. A codon-based model of nucleotide substitution for protein-coding DNA sequences. Mol. Biol. Evol. 11, 725–736 (1994)
Sanchez-Cespedes, M. et al. Inactivation of LKB1/STK11 is a common event in adenocarcinomas of the lung. Cancer Res. 62, 3659–3662 (2002)
Davies, H. et al. Mutations of the BRAF gene in human cancer. Nature 417, 949–954 (2002)
Granzier, H. L. & Labeit, S. Titin and its associated proteins: the third myofilament system of the sarcomere. Adv. Protein Chem. 71, 89–119 (2005)
Machado, C. & Andrew, D. J. D-Titin: a giant protein with dual roles in chromosomes and muscles. J. Cell Biol. 151, 639–652 (2000)
Machado, C., Sunkel, C. E. & Andrew, D. J. Human autoantibodies reveal Titin as a chromosomal protein. J. Cell Biol. 141, 321–333 (1998)
Zastrow, M. S., Flaherty, D. B., Benian, G. M. & Wilson, K. L. Nuclear Titin interacts with A- and B-type lamins in vitro and in vivo. J. Cell Sci. 119, 239–249 (2006)
Shiloh, Y. ATM and related protein kinases: safeguarding genome integrity. Nature Rev. Cancer 3, 155–168 (2003)
Renwick, A. et al. ATM mutations that cause ataxia-telangiectasia are breast cancer susceptibility alleles. Nature Genet. 38, 873–875 (2006)
Markowitz, S. et al. Inactivation of the type II TGF-beta receptor in colon cancer cells with microsatellite instability. Science 268, 1336–1338 (1995)
Howe, J. R. et al. Germline mutations of the gene encoding bone morphogenetic protein receptor 1A in juvenile polyposis. Nature Genet. 28, 184–187 (2001)
Wan, P. T. C. et al. Mechanism of activation of the RAF–ERK signaling pathway by oncogenic mutations of B-RAF. Cell 116, 855–867 (2004)
Irusta, P. M. et al. Definition of an inhibitory juxtamembrane WW-like domain in the platelet-derived growth factor beta receptor. J. Biol. Chem. 277, 38627–38634 (2002)
Teng, D. et al. Human mitogen-activated protein kinase kinase 4 as a candidate tumor suppressor. Cancer Res. 57, 4177–4182 (1997)
Su, G. et al. Alterations in pancreatic, biliary, and breast carcinomas support MKK4 as a genetically targeted tumor suppressor gene. Cancer Res. 58, 2339–2342 (1998)
Parsons, D. W. et al. Colorectal cancer Mutations in a signalling pathway. Nature 436, 792 (2005)
Bogoyevitch, M. A., Boehm, I., Oakley, A., Ketterman, A. J. & Barr, R. K. Targeting the JNK MAPK cascade for inhibition: basic science and therapeutic potential. Biochim. Biophys. Acta 1697, 89–101 (2004)
Kyriakis, J. M. & Avruch, J. Mammalian mitogen-activated protein kinase signal transduction pathways activated by stress and inflammation. Physiol. Rev. 81, 807–869 (2001)
Joshi-Tope, G. et al. Reactome: a knowledgebase of biological pathways. Nucleic Acids Res. 33, D428–D432 (2005)
Mi, H. et al. The PANTHER database of protein families, subfamilies, functions and pathways. Nucleic Acids Res. 33, D284–D288 (2005)
Kushida, T., Takagi, T. & Fukuda, K. Event ontology: a pathway-centric ontology for biological processes. Pac. Symp. Biocomput. 11, 152–163 (2006)
Wilkie, A., Patey, S., Kan, S., van den Ouweland, A. & Hamel, B. FGFs, their receptors, and human limb malformations: clinical and molecular correlations. Am. J. Med. Genet. 112, 266–278 (2002)
Cappellen, D. et al. Frequent activating mutations of FGFR3 in human bladder and cervix carcinomas. Nature Genet. 23, 18–20 (1999)
Sjoblom, T. et al. The consensus coding sequences of human breast and colorectal cancers. Science 314, 268–274 (2006)
Brose, M. S. et al. BRAF and RAS mutations in human lung cancer and melanoma. Cancer Res. 62, 6997–7000 (2002)
Acknowledgements
We would like to thank J. Leary and the ABN-Oncology group (funded by the National Health and Medical Research Council of Australia), the Hauenstein Foundation and the Cooperative Human Tissue Network for providing samples for analysis, G. Wu and L. Stein for the development of the joint Reactome, Panther, INOH database, and C. Marshall and N. Rahman for comments. The studies were funded by the NIH and the Wellcome Trust.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
Reprints and permissions information is available at www.nature.com/reprints. The authors declare no competing financial interests.
Supplementary information
Supplementary Information
This file contains Supplementary Methods and Supplementary Tables 1-5 with Legends and Supplementary Figure 1. The Supplementary Tables show protein kinase genes in the screen (Table 1); somatic mutations indentified (Table 2); cancer samples analysed (Table 3); germline variants identified (Table 4) and kinase gene ranking by probability of carrying driver mutations (Table 5). The Supplementary Figure 1 illustrates mutation prevalence in individual cancers across cancer types (PDF 2632 kb)
Rights and permissions
About this article
Cite this article
Greenman, C., Stephens, P., Smith, R. et al. Patterns of somatic mutation in human cancer genomes. Nature 446, 153–158 (2007). https://doi.org/10.1038/nature05610
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nature05610
This article is cited by
-
Lack of shared neoantigens in prevalent mutations in cancer
Journal of Translational Medicine (2024)
-
Hit discovery of potential CDK8 inhibitors and analysis of amino acid mutations for cancer therapy through computer-aided drug discovery
BMC Chemistry (2024)
-
Gsw-fi: a GLM model incorporating shrinkage and double-weighted strategies for identifying cancer driver genes with functional impact
BMC Bioinformatics (2024)
-
Emerging mechanisms of the unfolded protein response in therapeutic resistance: from chemotherapy to Immunotherapy
Cell Communication and Signaling (2024)
-
T-cell lymphocytes’ aging clock: telomeres, telomerase and aging
Biogerontology (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.