Abstract
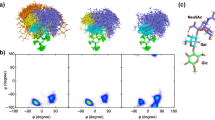
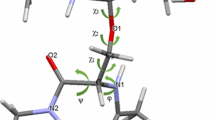
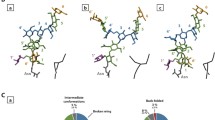
Glycosyltransferases acting onO-glycans have been shown to exhibit distinct specificity for the carbohydrate and the peptide moiety of their substrates. As an approach to study the 3-dimensional interactions between enzymes andO-glycan substrates, we determined the preferred conformations of five oligosaccharide-core structures of mucin type glycoproteins by NMR spectroscopy and by static and dynamic force field calculations. Seven oligosaccharides, representing five basic core structures, were investigated: Galβ(1–3)GalNAcαBzl (1, core 1), GlcNAcβ(1–6)[Galβ(1–3)]GalNAcαBzl (2, core 2), GlcNAcβ(1–3)GalNacαBzl (3, core 3), GlcNAcβ(1–6)[GlcNAcβ(1–3)]GalNAcαBzl (4, core 4), GlcNAcβ(1–6)GalNAcαBzl (5, core 6), the elongated core 2, Galβ(1–4)GlcNAcβ(1–6)[Galβ(1–3)]GalNAcαpNp (6) and GalNAcα-Bzl (7). The dynamic behaviour of the molecules was studied by Metropolis Monte Carlo (MMC) simulations. Experimental coupling constants, chemical shift changes, and NOEs were compared with results from static energy minimizations and dynamic MMC simulations and show a good agreement. MMC simulations show that the (1–6) linkage is much more flexible than the (1–3) or the (1–4) linkages. The preferred conformations of the disaccharides (1) and (3) show only slight differences due to the additionalN-acetyl group in (3). The conformational equilibrium of β(1–3) glycosidic bonds of1 and3 was not affected by attaching a β(1–6) linked GlcNAc unit to the GalNAc residue in2 and4. However, experimental and theoretical data show that the β(1–6) linkages of the trisaccharides2 and4, which carry an additional β(1–3) linked glycosyl residue, change their preferred conformations when compared with (5). The 6-branch also shows significant interactions with the benzyl aglycon altering the preferred conformation of the hydroxymethyl group of the GalNAc to a higher proportion of the gt conformer. The (1–6) linkage of2, 4, and6 can have two different families of conformations of which the lower energy state is populated only to about 20% of the time whereas the other state with a relative enthalpy of ≈4 kcal mol−1 is populated to 80%. This fact demonstrates that the two conformational states have different entropy contents. Entropy is implicitly included in MMC simulations but cannot be derived from energy minimizations.
Similar content being viewed by others
Abbreviations
- Bzl:
-
benzyl
- COSY:
-
correlation spectroscopy
- Gal:
-
d-galactose
- GalNAc:
-
N-acetyl-d-galactosamine
- GalNAc-ol:
-
N-acetylgalactosaminitol
- GlcNAc:
-
N-acetyl-d-glucosamine
- HOHAHA:
-
homonuclear Hartmann-Hahn-spectroscopy
- MMC:
-
metropolis Monte Carlo
- NOE:
-
nuclear Overhauser enhancement
- pNp:
-
p-nitrophenyl
- ROESY:
-
rotating frame Overhauser enhancement spectroscopy
- TOCSY:
-
totally correlated spectroscopy
References
Schachter H, Brockhausen I (1992) InGlycoconjugates, Composition, Structure and Function (Allen H, Kisailus EC, Eds), pp 263–332. New York: Marcel Dekker.
Brockhausen I, Möller G, Merz G, Adermann K, Paulsen H (1990)Biochemistry 29:10206.
Carver JP, Cummings D (1987)Pure Appl Chem 59:1465.
Lemieux RU, Bock K, Delbaere LTJ, Koto S, Rao SVR (1980)Can J Chem 58:631.
Thøgersen H, Lemieux RU, Bock K, Meyer B (1982)Can J Chem 60:44.
Duben AJ, Bush CA (1983)Arch Biochem Biophys 225:1.
Rao BN, Dua VK, Bush CA (1985)Biopolymers 24:2207.
Bush CA, Yan Z-Y, Rao BN (1986)J Am Chem Soc 108:6168.
Rao BN, Bush CA (1987)Biopolymers 26:1227.
Nishida Y, Hori H, Meguro H (1988)Agric Biol Chem 52:887.
Bechtel B, Wand AJ, Wroblewski K, Koprowski H, Thurin J (1990)J Biol Chem 265:2028.
Shogren R, Gerken TA, Jentoft N (1989)Biochemistry 28:5525.
Gerken TA, Jentoft N (1987)Biochemistry 26:4689.
Vliegenthart JFG, van Halbeek H, Dorland L (1981)Pure Appl Chem 53:70.
Klein A, Carnoy C, Lamblin G, Roussel P, van Kuik JA, de Waard P, Vliegenthart JFG (1991)Eur J Biochem 198:151.
Klein A, Carnoy C, Wieruszeski, Strecker G, Strang A-M, van Halbeek H, Roussel P, Lamblin G (1992)Biochemistry 31:6152.
Chai W, Hounsell E, Cashmore GC, Rosankiewicz JR, Feeney J, Lawson AM (1992)Eur J Biochem 207:973.
Chai W, Hounsell E, Cashmore GC, Rosankiewicz JR, Bauer CJ, Feeney J, Feizi T, Lawson AM (1992)Eur J Biochem 203:257.
Meyer B (1990)Topics Current Chem 154:141.
Poppe L, von der Lieth C-W, Dabrowski J (1990)J Am Chem Soc 112:7762.
Struike-Prill R, Meyer B (1990)Eur J Biochem 194:903.
Meyer B, Zsiska M, Struike-Prill R (1993)Computer Simulations in Condensed Matter Physics IV (Landau DP, Mon KB, Schuttler HB eds), Berlin: Springer-Verlag.
Brockhausen I, Williams D, Matta KL, Orr J, Schachter H (1983)Can J Biochem Cell Biol 61:1322.
Brockhausen I, Matta KL, Orr J, Schachter H (1985)Biochemistry 24:1866.
Brockhausen I, Matta KL, Orr J, Schachter H, Koenderman AHL, van den Eijnden DH (1986)Eur J Biochem 157:463.
Yazawa S, Abbas SA, Madiyalakan R, Barlow JJ, Matta KL (1986)Carbohydr Res 149:241.
Yousefi S, Higgins E, Doaling Z, Pollex-Krüger A, Hindsgaul O, Dennis J (1991)J Biol Chem 265:1772.
Flowers HM, Shapiro D (1965)J Org Chem 30:2041.
Piscorz CF, Abbas SA, Matta KL (1984)Carbohydr Res 126:115.
Abbas SA, Barlow JJ, Matta KL (1983)Carbohydr Res 112:201.
Abbas SA, Barlow JJ, Matta KL (1983)Carbohydr Res 113:63.
Kumar A, Wagner G, Ernst RR, Wüthrich K (1980)Biochem Biophys Res Commun 96:1156.
Davis DG, Bax A (1985)J Am Chem Soc 107:7197.
Summers MF, Marzilli LG, Bax A (1986)J Am Chem Soc 108:4285.
Bax A, Davis DG (1985)J Magn Reson 63:207.
Bax A, Davis DG (1985)J Magn Reson 65:355.
Metropolis N, Rosenbluth AW, Rosenbluth MN, Teller AH, Teller E (1953)J Chem. Phys 21:1087.
Peters T, Meyer B, Struike-Prill R, Somorjai R, Brisson J-R (1993)Carbohydr Res 238:49.
Fletcher R, Powell MJD (1963)Computer J 6:163.
Brisson J-R, Carver JP (1983)J Biol Chem 258:1431.
Bush CA, Feeney RE (1986)Int J Peptide Protein Res 28:386.
Ohrui H, Nishida Y, Hori H, Meguro H (1988)J Carbohydr Chem 7:711.
Neuhaus D, Keller J (1986)J Magn Reson 68:568.
Homans SW, DeVries AL, Parker SB (1985)FEBS Lett 183:133.
Maeji NJ, Inoue Y, Chujo R (1987)Int J Peptide Protein Res 29:699.
Gerlt JA, Joungblood VA (1980)J Am Chem Soc 102:7433.
Marchessault RH, Perez S (1979)Biopolymers 18:2369.
Nishida Y, Hori H, Ohrui H, Meguro H (1987)Carbohydr Res 170:106.
Ohrui H, Nishida Y, Higushi H, Hori H, Meguro H (1987)Can J Chem 65:1145.
Breg J, Kroon-Batenburg LMJ, Strecker G, Montreuil J, Vliegenthart JFG (1989)Eur J Biochem 178:727.
Streefkerk DG, De Bie MJA, Vliegenthart JFG (1973)Tetrahedron 29:833.
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Pollex-Krüger, A., Meyer, B., Stuike-Prill, R. et al. Preferred conformations and dynamics of five core structures of mucin typeO-glycans determined by NMR spectroscopy and force field calculations. Glycoconjugate J 10, 365–380 (1993). https://doi.org/10.1007/BF00731042
Received:
Issue Date:
DOI: https://doi.org/10.1007/BF00731042