Summary
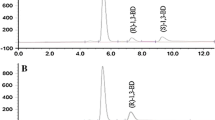
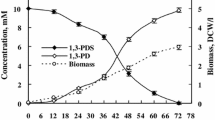
Thirty strains of microorganisms (bacteria, yeasts, fungi and green algae) were tested as resting free cells for their ability to transform acetyldimethylphenylsilane (1) enantioselectively into (R)-(1-hydroxyethyl)dimethylphenylsilane [(R)−2]. The biotransformations were monitored by GC (packed OV-17 column) and the enantiomeric purities of the products isolated were determined by HPLC (cellulose triacetate column, UV detection). All microorganisms tested were found to reduce 1 enantioselectively to give (R)-2. Under the test conditions used, the yeastTrigonopsis variabilis (DSM 70714) was found to exhibit the highest specific activity (1.5 mg product x g cell wet mass−1 x min−1), whereas the highest enantioselectivities were observed for the bacteriaAcinetobacter calcoaceticus (ATCC 31012) (>95% ee),Brevibacterium species (ATCC 21860) (90% ee) andCorynebacterium dioxydans (ATCC 21766) (>95% ee), the yeastCandida humicola (DSM 70067) (90% ee), the fungusCunninghamella elegans (ATCC 26269) (94% ee), as well as the cyanobacteriumSynechococcus leopoliensis (94% ee). From the green algae tested,Chlamydomonas reinhardii showed the highest enantioselectivity (85% ee).
Similar content being viewed by others
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Syldatk, C., Stoffregen, A., Wuttke, F. et al. Enantioselective reduction of acetyldimethylphenylsilane: A screening with thirty strains of microorganisms. Biotechnol Lett 10, 731–734 (1988). https://doi.org/10.1007/BF01025290
Issue Date:
DOI: https://doi.org/10.1007/BF01025290