Abstract
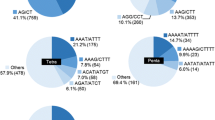
We have identified a set of informative microsatellite markers for genome analysis in kiwifruit and related Actinidia species. A small-insert genomic library was constructed from Actinidia chinensis DNA, and screened for microsatellites. About 1.2% of the total colonies hybridised to a (GA)8 probe, 0.4% to (GT)8, and 0.1% to a mixture of three different trinucleotide repeat probes, (CAA)5, (GAA)5 and (CTA)5. From the DNA sequences of 35 hybridising clones, 18 primer pairs were designed, and used to amplify genomic DNA from 38 individual plants, representing 30 different accessions of ten Actinidia species. The banding patterns for most of the dinucleotide repeats showed a high degree of polymorphism in the diploid and tetraploid A. chinensis, and in the hexaploid A. deliciosa (kiwifruit). Heterozygosity levels of up to 100% were found among eight diploid accessions of A. chinensis examined, and the number of different-sized bands among all the species varied from 3 to 36 for each microsatellite. One simple CT microsatellite gave 21 bands with sizes suggesting that the number of repeats ranged from 9 to 37. The highest number of bands (36) and the largest size variation (>100 bp) were observed with a complex microsatellite harbouring four different repeat motifs. The majority of primer pairs amplified bands from most of the ten Actinidia species tested. The most polymorphic primer pairs were used successfully to fingerprint a range of closely related varieties of kiwifruit (A. deliciosa).
Similar content being viewed by others
Abbreviations
- PCR:
-
polymerase chain reaction
- RFLP:
-
restriction fragment length polymorphism
- VNTR:
-
variable number of tandem repeats
References
Akkaya MS, Bhagwat AA, Cregan PB: Length polymorphisms of simple sequence repeat DNA in soybean. Genetics 132: 1131–1139 (1992).
Ali S, Müller CR, Epplen JT: DNA fingerprinting by oligonucleotide probes specific for simple repeats. Hum Genet 74: 239–243 (1986).
Atkinson RG: Molecular approaches to horticultural crop improvement. Ph.D. thesis, University of Auckland (1993).
Beckman JS, Soller M: Toward a unified approach to genetic mapping of eukaryotes based on sequence tagged microsatellite sites. Bio/technology 8: 930–932 (1990).
Bell CJ, Ecker JR: Assignment of 30 microsatellite loci to the linkage map of Arabidopsis. Genomics 19: 137–144 (1994).
Bierwerth S, Kahl G, Weigand F, Weising K: Oligonucleotide fingerprinting of plant and fungal genomes: A comparison of radioactive, colorigenic and chemiluminescent detection methods. Electrophoresis 13: 115–122 (1992).
Brunel D: A microsatellite marker in Helianthus annuus L. Plant Mol Biol 24: 397–400 (1994).
Clark JM: Novel non-templated nucleotide addition reactions catalyzed by prokaryotic and eukaryotic DNA polymerases. Nucl Acid Res 16: 9677–9685 (1988).
Condit R, Hubbell SP: Abundance and DNA sequence of two-base repeat regions in tropical tree genomes. Genome 34: 66–71 (1991).
Cregan PB, Bhagwat AA, Akkaya M, Rongwen J: Microsatellite fingerprinting and mapping of soybean. Meth Mol Cell Biol 5: 49–61 (1994).
Crowhurst RN, Gardner RC: A genome-specific repeat sequence from kiwifruit (Actinidia deliciosa var. deliciosa). Theor Appl Genet 81: 71–78 (1991).
Crowhurst RN, Lints R, Atkinson RG, Gardner RC: Restriction fragment length polymorphisms in the genus Actinidia (Actinidiaceae). Plant Syst Evol 172: 193–203 (1990).
Deka R, Shriver MD, Yu LM, Jin L, Aston CE, Chakraborty R, Ferrell RE: Conservation of human chromosome 13 polymorphic microsatellite (CA) n repeats in chimpanzees. Genomics 22: 226–230 (1994).
Devos KM, Bryan GJ, Collins AJ, Stephenson P, Gale MD: Application of two microsatellite sequences in wheat storage proteins as molecular markers. Theor Appl Genet 90: 247–252 (1995).
Edwards A, Civitello A, Hammond HA, Caskey CT: DNA typing and genetic mapping with trimeric and tetrameric tandem repeats. Am J Hum Genet 49: 746–756 (1991).
Ferguson AR: Kiwifruit: a botanical review. Hort Rev 6: 1–14 (1984).
Hearne CM, Ghosh S, Todd JA: Microsatellites for linkage analysis of genetic traits. Trends Genet 8: 288–294 (1992).
Janssen BJ, Gardner RC: The use of transient GUS expression to develop an Agrobacterium-mediated gene transfer system for kiwifruit. Plant Cell Rep 13: 28–31 (1993).
Kijas JMH, Fowler JCS, Garbett CA, Thomas MR: Enrichment of microsatellites from the Citrus genome using biotinylated oligonucleotide sequences bound to strepavidin-coated magnetic particles. Biotechniques 16: 656–662 (1994).
Kijas JMH, Fowler JCS, Thomas MR: An evaluation of sequence tagged microsatellite site markers for genetic analysis within Citrus and related species. Genome 38: 349–355 (1995).
Kimpton CP, Gill P, Walton A, Urquhart A, Millican ES, Adams M: Automated DNA profiling employing multiplex amplification of short tandem repeat loci. PCR Meth Appl 3: 13–22 (1993).
Lagercrantz U, Ellegren H, Andersson L: The abundance of various polymorphic microsatellite motifs differs between plants and vertebrates. Nucl Acids Res 21: 1111–1115 (1993).
Litt M, Hauge X, Sharma V: Shadow bands seen when typing polymorphic dinucleotide repeats: some causes and cures. Biotechniques 15: 280–284 (1993).
Moore SS, Sargeant LL, King TJ, Mattick JS, Georges M, Hetzel DJS: The conservation of dinucleotide microsatellites among mammalian genomes allows the use of heterologous PCR primer pairs in closely related species. Genomics 10: 654–660 (1991).
Morgante M, Olivieri AM: PCR-amplified microsatellites as markers in plant genetics. Plant J 3: 175–182 (1993).
Morgante M, Rafalski A, Biddle P, Tingey S, Olivieri AM: Genetic mapping and variability of seven soybean simple sequence repeat loci. Genome 37: 763–769 (1994).
Olson M, Hood L, Cantor C, Botstein D: A common language for physical mapping of the human genome. Science 245: 1434–1435 (1989).
Ostrander EA, Jong PM, Rine J, Duyk G: Construction of small-insert genomic DNA libraries highly enriched for microsatellite repeat sequences. Proc Natl Acad Sci USA 89: 3419–3423 (1992).
Pépin L, Amigues Y, Lépingle A, Berthier JL, Bensaid A, Vaiman D: Sequence conservation of microsatellites between Bos taurus (cattle), Capra hircus (goat) and related species. Examples of use in parentage testing and phylogeny analysis. Heredity 74: 53–61 (1995).
Röder M, Plaschke J, König SU, Börner A, Sorrells ME, Tanksley SD, Ganal M: Abundance, variability and chromosomal location of microsatellites in wheat. Mol Gen Genet 246: 327–333 (1995).
Rongwen J, Akkaya MS, Bhagwat AA, Lavi U, Cregan PB: The use of microsatellite DNA markers of soybean genotype identification. Theor Appl Genet 90: 43–48 (1995).
Saghai-Maroof MA, Biyashev RM, Yang GP, Zhang Q, Allard RW: Extraordinarily polymorphic microsatellite DNA in barley: species diversity, chromosomal locations, and population dynamics. Proc Natl Acad Sci USA 91: 5466–5470 (1994).
Sambrook J, Fritsch EF, Maniatis T: Molecular Cloning: A Laboratory Manual, 2nd ed. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, New York (1989).
Sanger F, Nicklen S, Coulson AR: DNA sequencing with chain-terminating inhibitors. Proc Natl Acad Sci USA 74: 5463–5467 (1977).
Schlötterer C, Amos B, Tautz D: Conservation of polymorphic simple sequence loci in cetacean species. Nature 254: 63–65 (1991).
Senior ML, Heun M: Mapping maize microsatellites and polymerase chain reaction confirmation of the targeted repeats using a CT primer. Genome 36: 884–889 (1993).
Smith DN, Devey ME: Occurrence and inheritance of microsatellites in Pinus radiata. Genome 37: 977–983 (1994).
Terauchi R: A polymorphic microsatellite marker from the tropical tree Dryobalanops lanceolata (Dipterocarpaceae). Jpn J Genet 69: 567–576 (1994).
Terauchi R, Konuma A: Microsatellite polymorphism in Dioscorea tokoro, a wild yam species. Genome 37: 794–801 (1994).
Thein SL, Wallace RB: The use of synthetic oligonucleotides as specific hybridization probes in the diagnosis of genetic disorders. In: Davies KE (ed) Human Genetic Diseases: A Practical Approach, pp. 33–50. IRL Press, Oxford (1986).
Thomas MR, Scott NS: Microsatellite repeats in grapevine reveal DNA polymorphisms when analysed as sequence-tagged sites. Theor Appl Genet 86: 985–990 (1993).
Thomas MR, Cain P, Scott NS: DNA typing of grapevines: A universal methodology and database for describing cultivars and evaluating genetic relatedness. Plant Mol Biol 25: 939–949 (1994).
Wang Z, Weber JL, Zhong G, Tanksley SD: Survey of plant short tandem repeats. Theor Appl Genet 88: 1–6 (1994).
Weber JL: Informativeness of human (dC-dA) n ×(dG-dT) n polymorphisms. Genomics 7: 524–530 (1990).
Weising K, Weigand F, Driesel AJ, Kahl G, Zischler H, Epplen JT: Polymorphic simple GATA/GACA repeats in plant genomes. Nucl Acids Res 17: 10128 (1989).
Weising K, Beyermann B, Ramser J, Kahl G: Plant DNA fingerprinting with radioactive and digoxigenated oligonucleotide probes complementary to simple repetitive DNA sequences. Electrophoresis 12: 159–169 (1991).
Wu KS, Tanksley SD: Abundance, polymorphism and genetic mapping of microsatellites in rice. Mol Gen Genet 241: 225–235 (1993).
Yan GJ, Ferguson AR, McNeilage MA: Ploidy in Actinidia chinensis. Euphytica 78: 175–183 (1994).
Yang GP, Saghai-Maroof MA, Xu CG, Zhang Q, Biyashev RM: Comparative analysis of microsatellite DNA polymorphism in landraces and cultivars of rice. Mol Gen Gent 245: 187–194 (1994).
Yu YG, Saghai-Maroof MA, Buss GR, Maughan PJ, Tolin SA: RFLP and microsatellite mapping of a gene for soybean mosaic virus resistance. Phytopathology 84: 60–64 (1994).
Zhao X, Kochert G: Phylogenetic distribution and genetic mapping of a (GGC)n microsatellite from rice (Oryza sativa L.). Plant Mol Biol 21: 607–614 (1993).
Zhou C, Yang Y, Jong AY: Mini-prep in ten minutes. Biotechniques 8: 172–173 (1990).
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Weising, K., Fung, R.W.M., Keeling, D.J. et al. Characterisation of microsatellites from Actinidia chinensis . Mol Breeding 2, 117–131 (1996). https://doi.org/10.1007/BF00441427
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/BF00441427