Abstract
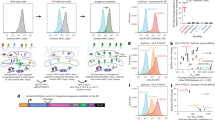
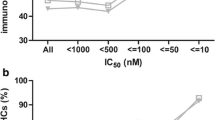
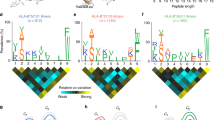
A peptide-based vaccine must be bound and presented by major histocompatibility complex class I molecules to elicit a CD8+ T-cell response. Because class I HLA molecules are highly polymorphic, it has yet to be established how well a vaccine peptide that stimulates one individual’s CD8+ cytotoxic T lymphocytes will be presented by a second individual’s different class I molecules. Therefore, to facilitate precise comparisons of class I peptide binding overlaps, we uniquely combined hollow-fiber bioreactors and mass spectrometry to assign precise peptide binding signatures to individual class I HLA molecules. In applying this strategy to HLA-B*1501, we isolated milligram quantities of B*1501-bound peptides and mapped them using mass spectrometry. Repeated analyses consistently assign the same peptide binding signature to B*1501; the degree of peptide binding overlap between any two class I molecules can thus be determined through comparison of their peptide signatures.
Similar content being viewed by others
Author information
Authors and Affiliations
Additional information
Received: 3 October 1996 / Revised: 20 November 1996
Rights and permissions
About this article
Cite this article
Prilliman, K., Lindsey, M., Zuo, Y. et al. Large-scale production of class I bound peptides: assigning a signature to HLA-B*1501. Immunogenetics 45, 379–385 (1997). https://doi.org/10.1007/s002510050219
Issue Date:
DOI: https://doi.org/10.1007/s002510050219