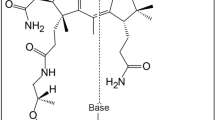
Abstract
SEVERAL hundred million tons of toxic mercurials are dispersed in the biosphere1. Microbes can detoxify organo-mercurials and mercury salts through sequential action of two enzymes, organomercury lyase2 and mercuric ion reductase (MerA) 3–5. The latter, a homodimer with homology to the FAD-dependent disulphide oxidoreductases6, catalyses the reaction NADPH + Hg(II) → NADP+ + H+Hg(0), one of the very rare enzymic reactions with metal substrates. Human glutathione reductase7,8 serves as a reference molecule for FAD-dependent disulphide reductases and between its primary structure9 and that of MerA from Tn501 (Pseudomonas), Tn21 (Shigella), pI258 (Staphylococcus) and Bacillus, 25–30% of the residues have been conserved10,11. All MerAs have a C-terminal extension about 15 residues long but have very varied N termini. Although the enzyme from Streptomyces lividans has no addition, from Pseudomonas aeruginosa Tn5Ol and Bacillus sp. strain RC607 it has one and two copies respectively of a domain of 80–85 residues, highly homologous to MerP, the periplasmic component of proteins encoded by the mer operon11. These domains can be proteolytically cleaved off without changing the catalytic efficiency3. We report here the crystal structure of MerA from the Gram-positive bacterium Bacillus sp. strain RC607. Analysis of its complexes with nicotinamide dinucleotide substrates and the inhibitor Cd(II) reveals how limited structural changes enable an enzyme to accept as substrate what used to be a dangerous inhibitor. Knowledge of the mode of mercury ligation is a prerequisite for understanding this unique detoxification mechanism.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Summers, A. O. & Silver, S. A. Rev. Microbiol. 32, 637–672 (1978).
Begley, T. P., Walts, A. E. & Walsh, C. T. Biochemistry 25, 7186–7192 (1986).
Fox, B. & Walsh, C. T. J. biol. Chem. 257, 2498–2503 (1982).
Misra, T. K. et al. Gene 34, 253–262 (1985).
Moore, M. J., Distefano, M. D., Zydowsky, L. D., Cummings, R. T. & Walsh, C. T. Acct chem. Res. 23, 301–308 (1990).
Williams, C. H. Jr The Enzymes 13, 90–165 (1976).
Karplus, P. A. & Schulz, G. E. J. molec. Biol. 195, 701–729 (1987).
Pai, E. F. & Schulz, G. E. J. biol. Chem. 258, 1752–1757 (1983).
Krauth-Siegel, R. L. et al. Eur. J. Biochem. 121, 259–267 (1982).
Laddaga, R. A., Chu, L., Misra, T. K. & Silver, S. Proc. natn. Acad. Sci. U.S.A. 84, 5106–5110 (1987).
Wang, Y. et al. J. Bacteriol. 171, 83–92 (1989).
Pai, E. F., Karplus, P. A. & Schulz, G. E. Biochemistry 27, 4465–4474 (1988).
Distefano, M. D., Au, K. G. & Walsh, C. T. Biochemistry 28, 1168–1183 (1989).
Distefano, M. D., Moore, M. J. & Walsh, C. T. Biochemistry 29, 2703–2713 (1990).
Miller, S. M. et al. Biochemistry 29, 2831–2841 (1990).
Miller, S. M., Moore, M. J., Massey, V., Williams, C. H. Jr & Walsh, C. T. Biochemistry 28, 1194–1205 (1989).
Miller, S. M., Ballou, D. P., Massey, V., Williams, C. H. Jr & Walsh, C. T. J. biol. Chem. 261, 8081–8084 (1986).
Miller, S. M., Massey, V., Williams, C. H. Jr, Ballou, D. P. & Walsh, C. T. Biochemistry 30, 2600–2612 (1991).
Rinderle, S. J., Booth, J. E. & Williams, J. W. Biochemistry 22, 869–876 (1983).
Distefano, M. D. thesis, Massachusetts Institute of Technology (1989).
Sahlmann, L., Lambeir, A.-M. & Lindskog, S. Eur. J. Biochem. 156, 479–488 (1986).
Raybuck, S. A., Distefano, M. D., Teo, B.-T., Orme-Johnson, W. & Walsh, C. T. J. Am. chem. Soc. 112, 1983–1989 (1990).
Moore, M. J., Distefano, M. D., Walsh, C. T., Schiering, N. & Pai, E. F. J. biol. Chem. 264, 14386–14388 (1989).
Kabsch, W. J. appl. Crystallogr. 21, 916–924 (1988).
Kabsch, W. J. appl. Crystallogr. 21, 67–71 (1988).
Dickerson, R. E., Weinzierl, J. E. & Palmer, R. A. Acta crystallogr. B24, 997–1003 (1968).
Jones, T. A. J. appl. Crystallogr. 11, 268–272 (1978).
Brünger, A. T., Kuriyan, J. & Karplus, M. Science 235, 458–460 (1987).
Read, R. J. Acta crystallogr. A42, 140–149 (1986).
Priestle, J. P. J. appl. Crystallogr. 21, 572–576 (1988).
Kabsch, W. & Sander, C. Biopolymers 22, 2577–2637 (1983).
Kabsch, W. Acta crystallogr. A34, 827–828 (1978).
Mattevi, A., Schierbeek, A. J. & Hol, W. G. J. J. molec. Biol. (in the press).
Kuriyan, J. et al. Nature 352, 172–174 (1991).
Karplus, P. A. & Schulz, G. E. J. molec. Biol. 210, 163–180 (1989)
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Schiering, N., Kabsch, W., Moore, M. et al. Structure of the detoxification catalyst mercuric ion reductase from Bacillus sp. strain RC607. Nature 352, 168–172 (1991). https://doi.org/10.1038/352168a0
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/352168a0
This article is cited by
-
Mechanisms of emerging resistance associated with non-antibiotic antimicrobial agents: a state-of-the-art review
The Journal of Antibiotics (2023)
-
Comparative genomics study reveals Red Sea Bacillus with characteristics associated with potential microbial cell factories (MCFs)
Scientific Reports (2019)
-
Modulation of the flavin–protein interactions in NADH peroxidase and mercuric ion reductase: a resonance Raman study
European Biophysics Journal (2018)
-
Structural and functional characterization of mercuric reductase from Lysinibacillus sphaericus strain G1
BioMetals (2017)
-
Structure of the membrane protein MerF, a bacterial mercury transporter, improved by the inclusion of chemical shift anisotropy constraints
Journal of Biomolecular NMR (2014)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.