-
PDF
- Split View
-
Views
-
Cite
Cite
Andrew L. Neal, Lani K. Clough, Todd D. Perkins, Brenda J. Little, Timothy S. Magnuson, In situ measurement of Fe(III) reduction activity of Geobacter pelophilus by simultaneous in situ RT-PCR and XPS analysis, FEMS Microbiology Ecology, Volume 49, Issue 1, July 2004, Pages 163–169, https://doi.org/10.1016/j.femsec.2004.03.014
Close - Share Icon Share
Abstract
Geobacter pelophilus is capable of dissimilatory Fe(III)-reduction on solid phase Fe(III)-oxides by means of surface attachment and direct electron transport to Fe(III), in part mediated by outer membrane c-type cytochromes. A study was undertaken to characterize surface colonization patterns, gene expression, and mineral transformation by this organism. The gene ferA (Geobacter sulfurreducens outer membrane Fe(III) reductase cytochrome c) was used as a target for PCR based molecular detection methods for visualizing G. pelophilus surface colonization. Protein extracts were prepared from solid-phase cultures, and cytochrome c content assessed. Mineral transformations were followed by X-ray photoelectron spectroscopy (XPS). Results of in situ (IS) RT-PCR experiments demonstrate that G. pelophilus attaches and grows at ferrihydrite mineral surfaces. Fluorescently-labeled cells were observed after IS-RT-PCR experiments, suggesting that G. pelophilus contains a cytochrome c sequence similar to ferA in G. sulfurreducens which is expressed in the presence of ferrihydrite. Protein extracts possessed high mass c-type cytochromes of similar size to those found in G. sulfurreducens. In addition, unique high-mass c-type cytochromes were also detected. XPS analysis demonstrated mineral transformation to occur, mediated by the surface associated population. This study demonstrates that G. pelophilus attaches to Fe(III)-oxide surfaces, reduces the Fe(III) oxides at the surface, produces c-type cytochromes under these growth conditions, and expresses cytochrome c-encoding genes as measured by in situ molecular detection techniques.
1 Introduction
Dissimilatory metal reduction is an important environmental process whereby microorganisms respire high valence metals and metal complexes. Ecologically, this process is significant in that it is responsible for many forms of mineral transformation in a variety of natural and contaminated environments. Dissimilatory metal-reducing bacteria (DMRB) have been found in many ecological niches; deep subsurface formations [1], freshwater and marine sediments [1], hydrothermal springs [2], and even in Antarctic ice [3]. The physiology and biochemistry of this process is increasingly well understood and can now be related to geochemical reactions catalyzed by this unique group of organisms. Research areas of emphasis include the biochemical analysis of redox proteins from metal-reducing organisms [4–7], genomic analysis of several representative organisms (Desulfovibrio gigas, Geobacter sulfurreducens, and Shewanella oniedensis), and the chemical reactions that are associated with respiratory metal reduction [8–14]. Much research involves the study of iron- and manganese-reducing organisms of the genera Geobacter and Shewanella. These are convenient model organisms for a variety of reasons; many representative isolates are available in pure culture, the organisms are relatively easy to culture on a variety of iron compounds, and the biochemistry and genetics of these organisms are increasingly well characterized.
While there has been much progress in the study of these organisms, major unresolved issues regarding the nature of mineral transformation remain, viz., the role of specific gene products (i.e., redox proteins) in the respiratory process, and whether these organisms do in fact attach to, and grow at mineral surfaces. While several noteworthy examples of research in these individual areas are in the literature [12,13,15–18], an integrated study utilizing both molecular and chemical analyses focused on mineral-based respiratory processes is lacking. The work presented here aims to address this issue, providing data linking gene expression with specific mineral transformation by a model organism.
Geobacter pelophilus is a dissimilatory metal-reducing bacterium closely related to the more studied G. sulfurreducens[19,20]. Although a great deal of knowledge continues to accumulate regarding Geobacter biochemistry, genetics, and physiology, few studies have been undertaken in order to study mineral surface associations and mineral transformation by other representatives of the Geobacteraceae. Using integrated techniques, a study was undertaken to characterize cell attachment to and gene expression and mineral transformation at ferrihydrite surfaces by this organism. The specific hypotheses to be tested were (1) G. pelophilus attaches to and modifies mineral surfaces; (2) G. pelophilus produces c-type cytochromes of similar physical properties as those of G. sulfurreducens and G. metallireducens; (3) G. pelophilus functional gene expression relevant to Fe(III) respiration can be measured at the single-cell level by means of in situ molecular techniques. X-ray photoelectron spectroscopy (XPS) was employed to take advantage of its surface specificity, localizing iron reduction at the mineral surface. XPS has clear advantages over wet-chemistry techniques where localization of reduction is not possible [21].
2 Materials and methods
2.1 Organism and culture conditions
Geobacter pelophilus [19,20] was obtained from the German Collection of Microorganisms and Cell Cultures (DSMZ). The organism was grown on modified freshwater media containing the following components (g/l): NH4Cl (1.5); NaH2PO4 (0.6); KCl (0.1); NaHCO3 (2.5); Wolfe's Vitamin Solution (10.0 ml); modified Wolfe's minerals (10.0 ml). All cultures were grown at 30 °C in anaerobic culture tubes under a 20% CO2/80% N2 atmosphere, and all culture manipulations were performed anaerobically. Acetate (20 mM) was provided as electron donor, and either Fe(III)-citrate (50 mM) or synthetic hydrous ferric oxide (HFO, ferrihydrite) (10% w/v) were provided as electron acceptor. Growth and cell division on surfaces was determined by microscopic monitoring (via total cell staining, see below) of cell counts over time, with maximal colonization observed after 5–7 days of incubation.
2.2 Preparation and analysis of DNA and RNA
Nucleic acids were prepared from log-phase cultures using commercial kits (QIAGEN). For preparation of RNA, cells were first treated with an RNAse inhibitor reagent prior to RNA preparation. Nucleic acids were quantified by UV spectrophotometry. Integrity of RNA preparations was determined by formaldehyde gel electrophoresis [22].
2.3 PCR and RT-PCR: ferA gene amplification
Sequences of G. sulfurreducens ferA (designated omcC[23]) and S. oniedensis MR-1 omcA were obtained from genome data at The Institute for Genome Research (http://www.tigr.org). Degenerate primers specific for the outer membrane cytochrome c (ferA) gene were designed from conserved regions (based on alignment of ferA with omcA) to target homologous genes in other Geobacter spp. Approximately 700 bp of the target was amplified. The primers used were OMC296F (5′-ACARMARSGRTGTYGGSTGC-3′) and OMC992R (5′-TGGATYMCSRGARGTYGYAGTG-3′). Careful design was performed to confirm that these primers recognized sequences only found in ferA and homologues. Primer specificity was confirmed by amplifying DNA or RNA from G. pelophilus. PCR was carried out under the following conditions: 200 nM forward and reverse primer; 20 mM Tris–HCl, pH 8.8; 50 mM KCl; 2.0 mM MgCl2; 1.0 U Taq DNA polymerase (Promega Corp., Madison, WI); 0.1% Tween20; and approximately 500 ng template DNA. Thermocycling began with a “hot-start” step of 5 min at 94 °C followed by 30 cycles of 94 °C for 30 s, 45 °C (ferA) for 1 min, and 72 °C for 2 min and ended with a 7 min 72 °C final extension. PCR products were analyzed by agarose gel (1.7%) electrophoresis, and visualized with ultraviolet light after ethidium bromide staining. Individual fragments with the predicted size were purified from gel slices with a QIAquick Gel Extraction kit (QIAGEN, Valencia, CA), according to the manufacturer's instructions. Products were cloned and plasmids extracted as above, and sequenced as described below using M13 forward and reverse primers. RT-PCR reactions contained 200 ng template RNA, 1 U of AMV-reverse transcriptase (Promega Madison, WI), 1 U of Tfl-DNA polymerase (Promega), 50 mM Tris–HCl, 63 mM KCl, 10 mM (NH4)2SO4, 5 mM dithiothrietol, and 10% (v/v) glycerol, 2 mM MgSO4, 0.2 mM dNTP, 200 μM each of forward and reverse primers, in a total reaction volume of 20 μl. Reverse transcription was carried out at 48 °C for 1 h, and after a 5 min RT inactivation step, PCR was continued using the reaction conditions described above. Products were analyzed as described above.
2.4 In situ RT-PCR
Cultures of G. pelophilus were grown on HFO or hematite containing media for 5 days, and the cells/mineral were then harvested by centrifugation, immediately fixed with 4% paraformaldehyde, washed three times with PBS, and treated with lysozyme (0.1 mg/ml) for 15 min to permeabilize the cells. Cells were washed again three times with PBS and then treated with DNase (5U/μl cell suspension) to digest DNA within the cells. The DNase was inactivated by heat, the cells washed, and resuspended in deionized water. This cell suspension was then used as template for in situ RT-PCR experiments. Primers OMC296F and OMC992R (specific for ferA) were used, and RT-PCR was carried out using AMV reverse transcriptase and Tfl DNA polymerase in a reaction mix that was optimized for in situ experiments. Reactions contained 5 μl cell suspension, 1 U of AMV-reverse transcriptase (Promega Madison, WI), 1 U of Tfl-DNA polymerase (Promega), 50 mM Tris–HCl, 63 mM KCl, 10 mM (NH4)2SO4, 5 mM dithiothrietol, 10% (v/v) glycerol, 2 mM MgSO4, 0.2 mM dNTP, 0.05 mM BODIPY-14-dUTP (Molecular Probes, Eugene, OR), and 200 nM each of forward and reverse primers, in a total reaction volume of 20 μl. Reactions were carried out in suspension in PCR tubes, with the same conditions listed above. After RT-PCR, cell suspension were washed three times with water and then mounted onto glass slides for microscopic observation. For total staining of all bacterial cells, RT-PCR treated specimens were stained with a solution of 4′,6-diamidino-2-phenylindole dihydrochloride (DAPI, Sigma Chemical Co., St. Louis, MO) in water for 10 min, followed by a brief destaining with deionized water.
2.5 Microscopy and image analysis
Microscopy analysis of bacterial cells to the solid phase mineral particles was performed using a Nikon Eclipse E800 microscope equipped with 40× and 60× objectives, DIC polarizing filters, and epifluorescence capability. For epifluorescence microscopy, samples were illuminated with a 100 W Hg-vapor discharge lamp, and fluorescence was detected and filtered by means of UV-2A (DAPI) or HYQ Cy5 (Cy5) filter cubes (Nikon). Image capture was performed using a Micromax RTE/CCD-732–7 (Princeton Instruments, Trenton, NJ) camera and MetaVue 5.0 software.
2.6 Protein extraction and analysis
G. pelophilus was grown on minimal medium (described above) containing 10% (w/v) HFO as electron acceptor, and after sufficient colonization had developed, HFO was harvested by centrifugation at 5000g for 15 min. The solids were washed twice in 50 mM Tris–HCl buffer and resuspended in the same buffer. HFO suspensions were treated with lysozyme (0.1 mg/ml final concentration) for 30 min at room temperature, and then SDS was added to 1% final concentration. The HFO was then extracted with gentle mixing for 1 h at room temperature. Solids and insoluble material was removed by centrifugation (10,000g, 10 min), and the resulting supernatant was concentrated approximately 100-fold by ultrafiltration. The extracts were then resolved on 10% SDS–PAGE gels [24], and stained for hemoprotein (cytochrome c) [25] and total protein content [24].
2.7 X-ray photoelectron spectroscopy
X-ray photoelectron spectroscopy (XPS) was employed to determine the valence state of Fe, and thus the extent of iron reduction, at both uncolonized and colonized iron oxide surfaces. XPS is a surface sensitive (penetration depth ∼100 Å) spectroscopic technique ideally suited to identification of chemical transformations at surfaces. Spectra were collected on a Perkin–Elmer Physical Electronics Division Model 5600ci spectrometer (Perkin–Elmer Inc., Eden Prairie, MN). The spectrometer was calibrated employing the Au 4f7/2, Cu 2p3/2 and Ag 3d5/2 photopeaks with binding energies of 83.99, 932.66 and 368.27 eV, respectively. A consistent 400 μm spot size was analyzed on all surfaces using a monochromatized Al Kα (hν= 1486.6 eV) X-ray source at 300 W and a pass energy of 46.95 eV for survey scans, and 11.75 eV for high-resolution scans. The system was operated at a base pressure of 10−8 to 10−9τ. A consistent emission angle (2φ) of 45° was used throughout. Following baseline subtraction [26], curves were fit employing an 80% Gaussian: 20% Lorenztian line shape. A common problem associated with the analysis of insulating materials such as iron oxides is the accumulation of surface charge during spectral collection leading to photopeak shifts. This potential problem was overcome by the use of a 5 eV flood gun and by referencing of the principal C 1s photopeak (nominally due to carbon of the type [–CH2–CH2–]n) to a binding energy (Eb) of 284.8 eV [27]. Ferrihydrite samples were dried onto a Si wafer before mounting for XPS analysis. Samples were mounted in preparation for XPS in an anaerobic glove box and transported to the spectrometer in a plate chamber sealed under an anaerobic atmosphere. Brief (<5 s) exposure to air was unavoidable during introduction of the samples into a N2 flushed antechamber. This antechamber was then evacuated and the sample placed into the spectrometer itself.
2.8 Analysis of Fe valence state
In the transition metals, unpaired electrons give rise to so-called multiplet states. These states may have widely differing Eb and may thus be represented in photoelectron spectra by different photopeaks. Any model capable of differentiating between Fe(III) and Fe(II) ions must take into account these multiplet states. Theoretical core p level models describing multiplet splitting associated with ferrous and ferric ions have been demonstrated by Gupta and Sen [28] and empirical proofs substantiated by Pratt et al. [29]. Accordingly, here we fit ferric ion contributions to high-resolution Fe 2p3/2 core regions with five peaks that decrease in intensity at increased Eb, all peaks having the same line-shape. Ferrous ion contributions are represented by a major peak accompanied by a pair of multiplet peaks 0.9 eV either side of the major peak and a shake-up satellite at elevated Eb (ΔEb∼6 eV, [30]). The relative intensities of the Fe(II) peaks are consistent with the aforementioned models and have been applied elsewhere to good effect in the identification of reduced iron in the presence of microorganisms [14,31].
3 Results
3.1 Population level gene expression patterns based on RT-PCR
Biochemical evidence shows that both G. sulfurreducens and G. pelophilus synthesize a variety of c-type cytochromes, presumably serving as electron transport mediators in Fe(III) respiration [5] Using degenerate primers targeting ferA from G. sulfurreducens, an amplicon was generated from G. pelophilus DNA that encoded a cytochrome sequence. Analysis of RNA extracted from cultures of G. pelophilus grown on hydrous ferric oxide (HFO) demonstrates expression of this gene, with a single band of correct size observed on agarose gels.
3.2 Gene expression on mineral surfaces based on in situ PCR
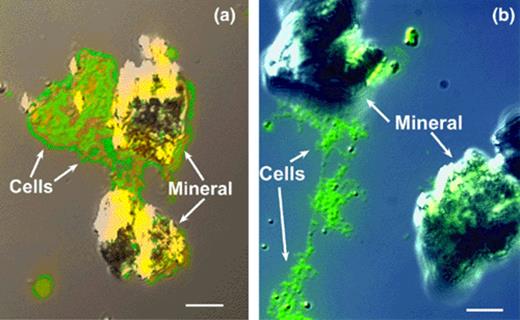
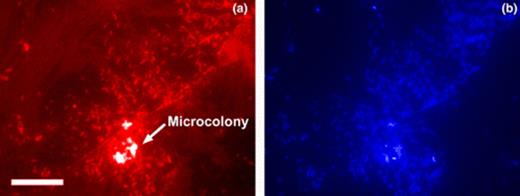
When G. pelophilus is grown in the presence of HFO, cells associate with and colonize the mineral surface (Fig. 1(a)). Differential interference contrast (DIC) images also reveal polymeric material enveloping the microcolonies (Fig. 1(a) and (b)). Cells cultured in the presence of hematite show a similar pattern, with clear labeling of cells on the hematite surface (Fig. 2). DAPI stained images (Fig. 2(b)) give an indication of gene-specific labeling efficiency, and serves to further illustrate surface colonization by G. pelophilus.
(a) and (b). In situ RT-PCR analysis and DIC microscopy of ferA gene expression and surface-associated growth on hydrous ferric oxide (HFO) surface by G. pelophilus attached to HFO particles. The fluorescence image was overlayed onto the DIC image. Panel a, tightly associated biofilm attached to mineral surface; panel b, tightly associated and detached biofilm. Bar equals 10 μm.
G. pelophilus attachment to hematite surface as visualized by in situ RT-PCR and total cell staining. Panel a, PCR labeled cells with ferA as target; panel b, DAPI stained cells. Note presence of high cell density ‘microcolony’ on surface. Bar equals 10 μm.
3.3 Cytochrome composition
Polyacrylamide gel elctrophoresis experiments demonstrate that both G. sulfurreducens and G. pelophilus produce suites of c-type cytochromes, ranging in mass from 100 to 10 kDa (Data not shown). This pattern is also observed with other members of the Geobacter genus, including G. bremensis and G. grbiciae (Magnuson, unpublished data).
3.4 Spectroscopic analysis of bacteria-free and bacterial-colonized iron oxide surfaces
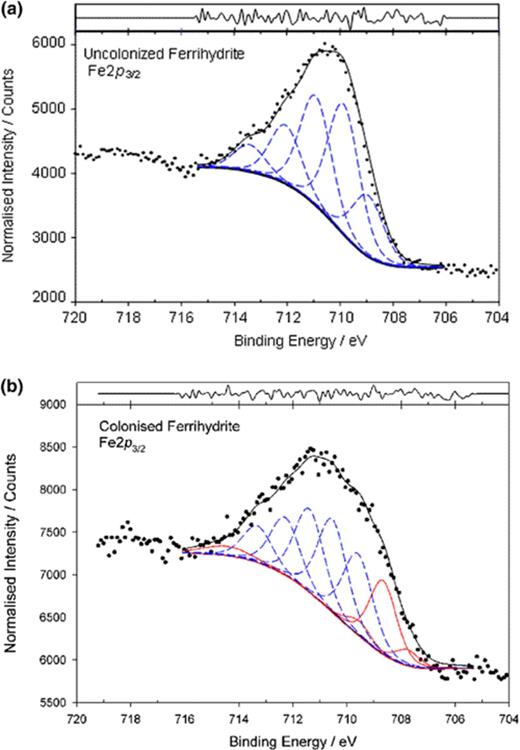
High-resolution core Fe 2p3/2 spectra collected on bacteria-free ferrihydrite surfaces exposed to culture medium suggested no abiotic Fe-reduction took place during incubation (Fig. 3(a)). The Fe(III) multiplet splitting model provided an adequate fit to the peak envelope on both surfaces and is in agreement with published iron oxide spectra [8,30]. However, following colonization by G. pelophilus the Fe(III) model no longer provided an adequate description of the observed peak envelope. Addition of a principal Fe(II) peak at 708.4 eV for the ferrihydrite spectra, (together with associated multiplet and satellite peaks), was required to complete the model (Fig. 3(b)). These results suggest that Fe-reduction occurred on those ferrihydrite surfaces colonized by G. pelophilus, a contribution of 53.3 at.% Fe(II) was identified on ferrihydrite (Fig. 3). In contrast, no detectable conversion of hematite was observed.
XPS spectra obtained for uncolonized (a) and colonized (b) ferrihydrite surfaces. Blue lines represent Fe(III) multiplet peaks; red lines represent Fe(II) multiplet peak assignments. C 1s=284.8 eV.
4 Discussion
The process of dissimilatory Fe(III) reduction has typically been studied from either a microbiological or geochemical perspective. It is becoming increasingly apparent however, that use of complementary molecular biological, biochemical, and geochemical approaches offers the opportunity to obtain a more comprehensive understanding of the overall biogeochemical process. The results of the present study describe the intimate relationship between cells of G. pelophilus expressing cytochrome c genes on Fe-mineral surfaces and dissimilatory Fe-reduction. Biochemical evidence suggests that at least one membrane-associated cytochrome c with Fe(III)-reductase activity is expressed at higher levels during growth on soluble Fe(III) [5,32]. This evidence is useful in choosing potential target genes for in situ techniques involving mineral colonization by dissimilatory metal reducing bacteria (DMRB). Previous studies have implicated both membrane-associated cytochromes and hydrogenases in reduction of soluble phase metals in both DMRB and SRB [4–6,9,11].
Protein analysis indicates that both G. pelophilus and G. sulfurreducens produce a suite of c-type cytochromes when grown with HFO as terminal electron acceptor. FerA, the 89 kDa cytochrome c from G. sulfurreducens characterized by Magnuson et al. [32], is produced under these conditions, as well as 78 and 36 kDa cytochromes. Recent work by Leang et al. [23] has shown that both FerA (OmcC) and the 78 kDa cytochrome c (OmcB) cytochromes were expressed under growth with soluble Fe(III), but no data was presented that showed production of these cytochromes with solid-phase minerals. Our work with both G. sulfurreducens and G. pelophilus presented here clarifies this issue. G. pelophilus similarly produces several cytochromes c among these a high-mass cytochrome of approximately 110 kDa. G. metallireducens also produces a >100 kDa cytochrome c under Fe(III)-grown conditions (Magnuson, unpublished data). In S. oniedensis MR-1, it has been shown that outer membrane cytochromes are exposed to the exterior, and thus able to interact with solid-phase electron acceptors [33]. From a comparative biochemistry standpoint, it is clear that all of these organisms produce high-mass cytochromes, presumably similar in function, as key components of their electron transport chains. This is the first report of comparative cytochrome profiling with Geobacter spp. grown with solid-phase electron acceptors. We are currently characterizing and comparing cytochrome profiles and individual high-mass cytochromes from several additional representatives of the Geobacteraceae in order to better understand their function
It can be concluded that attached populations of G. pelophilus significantly alter the chemistry of amorphous Fe(III) oxides. X-ray photoelectron spectroscopy provides a useful alternative to conventional wet-bench chemical measurements. Using this technique, it is possible to analyze mineral samples on the same scale and from the same culture as those used for molecular analysis. Surface analysis of G. pelophilus colonized minerals provides strong evidence for bacterial transformation and alteration of the surface chemistry of amorphous Fe(III) oxides. As was observed for G. sulfurreducens, G. pelophilus also attaches to and modifies Fe-oxide surfaces. Childers et al. [15] presented evidence for direct interaction of G. metallireducens with Fe(III) oxide surfaces, and proposed that cell wall proteins (pilins, flagella) were involved in adhesion to these surfaces. However, no surface analysis was performed to demonstrate mineral transformation. Studies by Das and Caccavo [17,18], using Shewanella alga have also demonstrated this phenomenon, and some intriguing biochemical evidence was presented for an attachment mechanism. However, that work did not suggest specific gene expression patterns. Our results clearly demonstrate the utility of in situ expression and protein synthesis analysis in the characterization of surface colonization by Geobacter spp. With these methods, direct comparison of different DMRB is possible, and in the case of Geobacter spp., some similar features are observed. First, these organisms all produce multiple high-mass cytochromes. Second, Geobacter spp. appear to be capable of significant conversion of ferrihydrite, although some strains (G. pelophilus in particular) may not have the ability to transform more crystalline forms of Fe(III)-oxides (e.g., hematite). The data presented here clearly manifest that G. pelophilus attaches to both Fe(III) oxide and oxyhydroxide studied, and probably has an affinity high enough to avoid detachment during in situ analysis. In addition, it is demonstrated that molecular techniques are employable in systems containing bacteria and Fe(III) oxides. The cells establish an interaction with these surfaces that enable at least some of them to avoid displacement from the mineral surface during preparation for in situ analysis.
The study also demonstrates the use of in situ molecular and biochemical techniques in systems containing bacteria and Fe(III) oxides. Although preliminary investigations suggested RT-PCR assay interference by the Fe mineral phases (possibly the result of RT-PCR enzymes, primers, and labels adsorption to the mineral phase), assay conditions were identified that enabled detection of gene expression in cells in intimate contact with these mineral surfaces. In addition, protein extracts can be obtained from the colonized material in sufficient quantity for biochemical analysis. It can be concluded, therefore, that surface analysis and molecular/biochemical techniques can be combined to detect and relate specific surface-associated mineral biotransformations to the expression of specific functional genes in individual cells and microcolonies on the mineral surface. These techniques are being employed in this integrated manner to characterize mineral colonization and biotransformation, and the genes involved in these processes by other members of the Geobacteraceae. It is anticipated that these techniques will also be applicable to ecological studies where in situ gene detection and mineral transformation are of interest.
Acknowledgements
XPS was performed at the Image and Chemical Analysis Laboratory, Department of Physics, Montana State University, Bozeman. Funding was provided by the Office of Naval Research, the US Department of Energy-Natural and Accelerated Bioremediation Research Program, Idaho State University and NSF-EPSCoR start-up funds (to T.S.M.), and the University of Georgia Research Foundation (to A.L.N.).
References